 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OPUNC01G08460.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 799aa MW: 86498 Da PI: 9.5075 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 38.9 | 1.6e-12 | 645 | 691 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
OPUNC01G08460.1 645 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITYITDLQ 691
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 8.3E-50 | 241 | 426 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.831 | 641 | 690 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.44E-18 | 644 | 708 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.52E-15 | 644 | 695 | No hit | No description |
| Pfam | PF00010 | 5.2E-10 | 645 | 691 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 9.1E-18 | 645 | 704 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-16 | 647 | 696 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 799 aa Download sequence Send to blast |
MLVEHATTLY ESPSREEPRG RRSSRLLHPP PPNEPRLVSC RLGSGSNPTG VEKARPPSFS 60 FHTSPPPPLS PHHKSPPSSA PEIPSRTRET RGTTTTTTTA SCARGEFGDP RASVGGGGEI 120 RASPSPGVLG FSHQRRGEGG ARLFFSRGAG KERGPVRPGF RRRGADWGAG APARWGLRRE 180 EEEGRMEVDE DGANGGTGGT WTDEDRALSA SVLGTDAFAY LTKGGGAISE GLVAASLPVD 240 LQNRLQELVE SDRPGAGWNY AIFWQLSRTK SGDLVLGWGD GSCREPRDGE MGPAASAGSD 300 EAKQRMRKRV LQRLHSAFGG VDEEDYAPGI DQVTDTEMFF LASMYFAFPR RAGGPGQVFA 360 AGVPLWIPNT ERNVFPANYC YRGYLANAAG FRTIVLVPFE SGVLELGSMQ QVAESSDTLQ 420 TIRSVFAGAI GNKAGVQRHE GNGPADKSPG LAKIFGKDLN LGRPSAGPGT GVSKADERSW 480 EQRTAGGGSS LLPNVQRGLQ NFTWSQARGL NSHQQKFGNG ILIVSNEATP RNNGVVDSPT 540 ATQFQLQKAS PLQKLPQLQK PHQIVKPQQL VSQQQLQPQA PRQIDFSAGT SSKPGVLTKK 600 PAGIDGEGAE VDGLCKDEGP PPAIEDRRPR KRGRKPANGR EEPLNHVEAE RQRREKLNQR 660 FYALRAVVPN ISKMDKASLL GDAITYITDL QKKLKEMEVE RERLIESGMV DPRERTPRPE 720 VDIQVVQDEV LVRVMSPMEN HPVRAIFQAF EEAEVHAGES KITSNNGTAV HSFIIKCPGA 780 EQQTREKVIA AMSRAMNSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 7e-25 | 241 | 426 | 9 | 192 | Transcription factor MYC3 |
| 4rqw_B | 7e-25 | 241 | 426 | 9 | 192 | Transcription factor MYC3 |
| 4rs9_A | 7e-25 | 241 | 426 | 9 | 192 | Transcription factor MYC3 |
| 4yz6_A | 7e-25 | 241 | 426 | 9 | 192 | Transcription factor MYC3 |
| 5gnj_A | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 3e-26 | 639 | 701 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 626 | 634 | RRPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
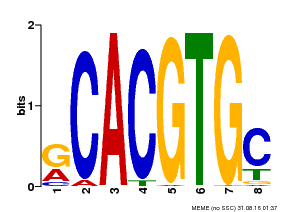
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012612 | 0.0 | CP012612.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 4 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015621308.1 | 0.0 | transcription factor bHLH13 | ||||
| TrEMBL | A0A0E0JG39 | 0.0 | A0A0E0JG39_ORYPU; Uncharacterized protein | ||||
| STRING | OPUNC01G08460.1 | 0.0 | (Oryza punctata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-69 | bHLH family protein | ||||



