 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | ONIVA02G06230.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1035aa MW: 116220 Da PI: 5.8541 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.7 | 3e-55 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
ke ++rwl+++ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+een +fq
ONIVA02G06230.1 20 KEaQRRWLRPTEICEILKNYRSFRIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKSGSIDVLHCYYAHGEENINFQ 113
55599***************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+Lee++ +ivlvhylevk
ONIVA02G06230.1 114 RRSYWMLEEDYMHIVLVHYLEVK 136
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.606 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 5.2E-76 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.6E-48 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.2E-7 | 446 | 531 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 0.00346 | 448 | 535 | No hit | No description |
| SuperFamily | SSF81296 | 1.57E-17 | 448 | 533 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.6E-8 | 448 | 523 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 8.86E-17 | 637 | 741 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 4.8E-18 | 639 | 745 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.21E-13 | 644 | 738 | No hit | No description |
| PROSITE profile | PS50088 | 8.763 | 646 | 678 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 18.059 | 646 | 750 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.013 | 679 | 708 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.843 | 679 | 711 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1400 | 718 | 747 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 4.78E-8 | 850 | 903 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.075 | 852 | 874 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.407 | 853 | 882 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0034 | 854 | 873 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 875 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 876 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 878 | 897 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1035 aa Download sequence Send to blast |
MAEGRRYAIA PQLDIEQILK EAQRRWLRPT EICEILKNYR SFRIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKSGSIDVLH CYYAHGEENI NFQRRSYWML 120 EEDYMHIVLV HYLEVKLKFH FLHIYGYDHQ AGKLSSRSTG HDDVLQVSHA DSPLSQLPSQ 180 TTEGESSVSG QASEYDETES AYLKQNSVQQ IFIREEPDTT LSLGCGSMKM EVDLYQGLQA 240 TAPNTGFYSH GQDNLPVVLN ESDLGTAFNG PNSQFDLSLW IEAMKPDKGT HQIPLYQAPV 300 PSEQSPFTGG PGIESFTFDE VYNNGLSIKD VDGDDTDGET PWQIPNASGT FATADSFQQN 360 DKTLEEAINY PLLKTQSSSL SDIIKDSFKK NDSFTRWMSK ELAEVDDSQI TSSSGVYWNS 420 EEADNIIEAS SSDQYTLGPV LAQDQLFTIV DFSPTWTYAG SKTRVFIKGN FLSSDEVKRL 480 KWSCMFGEFE VPAEIIADDT LVCHSPSHKP GRVPFYVTCS NRLACSEVRE FDFRPQYMDA 540 PSPLGSTNKI YLQKRLDKLL SVEQDEIQTT LSNPTKEIID LSKKISSLMM NNDDWSELLK 600 LADDNEPATD DKQDQFLQNR IKEKLHIWLL HKVGDGGKGP SMLDEEGQGV LHLAAALGYD 660 WAIRPTIAAG VNINFRDAHG WTALHWAAFC GRERTVVALI ALGAAPGAVT DPTPSFPSGS 720 TPADLASANG HKGISGFLAE SSLTSHLQTL NLKEAMRSSA GEISGLPGIV NVADRSASPL 780 AVEGHQTGSM GDSLGAVRNA AQAAARIYQV FRMQSFQRKQ AVQYEDENGA ISDERAMSLL 840 SAKPSKPAQL DPLHAAATRI QNKFRGWKGR KEFLLIRQRI VKIQAHVRGH QVRKHYRKII 900 WSVGIVEKVI LRWRRRGAGL RGFRPTENAV TESTSSSSGN VTQNRPAEND YDFLQEGRKQ 960 TEERLQKALA RVKSMVQYPD ARDQYQRILT VVTKMQESQA MQEKMLEEST EMDEGLLMSE 1020 FKELWDDDMP TPGYF |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
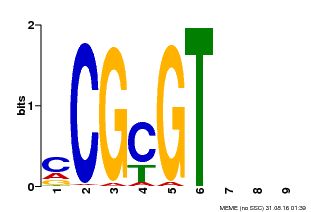
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK066651 | 0.0 | AK066651.1 Oryza sativa Japonica Group cDNA clone:J013074G10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015631248.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X1 | ||||
| Refseq | XP_015631249.1 | 0.0 | calmodulin-binding transcription activator 1 isoform X2 | ||||
| TrEMBL | A0A0E0G277 | 0.0 | A0A0E0G277_ORYNI; Uncharacterized protein | ||||
| STRING | ONIVA02G06230.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



