 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART09G15750.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 407aa MW: 43712.4 Da PI: 7.479 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.7 | 1.1e-40 | 45 | 121 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC+adls+ ++yhrrhkvCe hsk+pvv+vsg+e rfCqqCsrfh l efDe+krsCr+rL++hn+rrrk+qa
OBART09G15750.1 45 CAVDGCKADLSKHRDYHRRHKVCEPHSKTPVVVVSGREMRFCQQCSRFHLLGEFDEAKRSCRKRLDGHNRRRRKPQA 121
**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 2.6E-36 | 37 | 106 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.923 | 42 | 119 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.83E-38 | 43 | 124 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 9.9E-32 | 45 | 118 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 407 aa Download sequence Send to blast |
MDWDLKMPVS WDLAELEHNA VPPSTSTLKR PRGGGGGGGG QCPSCAVDGC KADLSKHRDY 60 HRRHKVCEPH SKTPVVVVSG REMRFCQQCS RFHLLGEFDE AKRSCRKRLD GHNRRRRKPQ 120 ADSMSSGSFM TSQQGTRFAS FTPPRPEPSW PGIIKSEETP YYSHHHHPHP VMTSRQPHFV 180 GSPSSATTAA FSPKEGRRFP FLHEGDQISF GGGGGAAAAA TLEISVCQPL LKTTVVAPPP 240 PESSSSNKMF SSDGLTTATT TTTTAHHHHH HHQVLDSDCA LSLLSSPANS SSVDPDGPAV 300 TGGGAGAEHH HHHHHHQIPM AQPLVPNLQQ QFGGSSPWFA SSPAAAAVAG GGFACPSMDS 360 EQQQQQQLNA VLVPGSNENE MNYHGMFHVG GEGSSDGTSP SLPFSWQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-27 | 35 | 118 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 101 | 118 | KRSCRKRLDGHNRRRRKP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
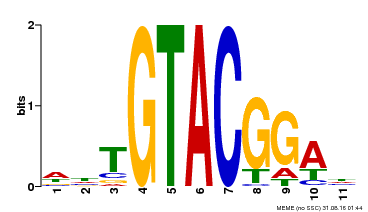
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by microRNAs miR156b and miR156h. {ECO:0000305|PubMed:16861571}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC108759 | 0.0 | AC108759.2 Oryza sativa (Japonica cultiva-group) chromosome 9 BAC clone OSJNBa0072P02, complete sequence. | |||
| GenBank | AP014965 | 0.0 | AP014965.1 Oryza sativa Japonica Group DNA, chromosome 9, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025876170.1 | 0.0 | squamosa promoter-binding-like protein 18 isoform X3 | ||||
| Swissprot | Q0J0K1 | 0.0 | SPL18_ORYSJ; Squamosa promoter-binding-like protein 18 | ||||
| TrEMBL | A0A0D3H8P6 | 0.0 | A0A0D3H8P6_9ORYZ; Uncharacterized protein | ||||
| STRING | OBART09G15750.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1755 | 37 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 1e-39 | SBP family protein | ||||



