 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OBART01G31160.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 786aa MW: 86298.4 Da PI: 8.9863 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 78.3 | 8e-25 | 191 | 292 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr 87
f+k+lt sd++++g +++p++ ae++ ++ s++l+ +d++g +W++++iyr++++r++lt+GW++Fv+ ++L +gD v+F +
OBART01G31160.1 191 FCKTLTASDTSTHGGFSVPRRAAEDCfppldhkQLR--PSQELVAKDLHGAKWRFRHIYRGQPRRHLLTTGWSSFVNKKKLVSGDAVLFL--RG 280
99**************************99865444..4669************************************************..34 PP
SEE..EEEEE-S CS
B3 88 sefelvvkvfrk 99
+++el+++v+r+
OBART01G31160.1 281 DDGELRLGVRRA 292
89999****997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.14E-47 | 175 | 330 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 6.8E-39 | 186 | 294 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.12E-22 | 189 | 291 | No hit | No description |
| Pfam | PF02362 | 7.8E-23 | 191 | 292 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.507 | 191 | 293 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.8E-24 | 191 | 293 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.7E-31 | 318 | 397 | IPR010525 | Auxin response factor |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010050 | Biological Process | vegetative phase change | ||||
| GO:0010158 | Biological Process | abaxial cell fate specification | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 786 aa Download sequence Send to blast |
MASSASSSSS PSSRPPLMAL PSFYRPPWPS ERGGEQRATD CWAGSPAAGG GRARATAMGI 60 DLNNTASGGE EDAPAPAPVC RDLWHACAGP VVSLPRRGSA VVYLPQGHLS AAGAGGGIRG 120 EVAVALPPHV ACRVVDVELC ADAATDEVYA RLALRAEGEV FERNLHGGGI EREDDMEDGD 180 EERKSRMLHM FCKTLTASDT STHGGFSVPR RAAEDCFPPL DHKQLRPSQE LVAKDLHGAK 240 WRFRHIYRGQ PRRHLLTTGW SSFVNKKKLV SGDAVLFLRG DDGELRLGVR RATQLKNEAI 300 FKAFSSESSK MRTLSAVADS LKHGSVFHIC YNPRATASEY VVPYWKFVKS FNHPVCIGMR 360 FKFHFESEDV NERRSGMIAG VSEVDPIRWP GSKWRSLLVR WEDATDCNSQ NRVSPWEIEI 420 VGGSISVAHS LSASNGNGRP DSVETEKFPR VLQGQELMGS RTHRVTCSPQ SIDITKSKSF 480 DAWRFLTDTR SCMLGSSTSR LPVQYSGYTH QSVSFGESIG FPEVLQGQEI SQTVPPFQGM 540 LPDACSAKSR YELKNYVCTP ATMNGLSSAN EGYCLSLSTV PPSPPSSLML YQTGVPQLEL 600 ASKNNDKSGN DSQPALRQHK LLSETSWDQF KIGKASTPGN ATKPGNGGRE VDRTSSRLFG 660 FSLTEKIIPT DKDGEKEVSY ETDCQNPRML DLFGYNCSTP EVNARFARCT SGARRGKSLI 720 TSVGSSRYVN QIMHTAAYLF SKESWNGWYC RIVKGSAHRP KLKLYHPKPI VILSRVLKNH 780 FYGKSK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldy_A | 1e-100 | 79 | 419 | 19 | 353 | Auxin response factor 1 |
| 4ldy_B | 1e-100 | 79 | 419 | 19 | 353 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
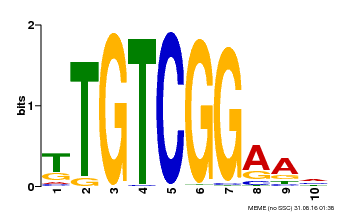
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00021 | PBM | Transfer from AT2G33860 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072330 | 0.0 | AK072330.1 Oryza sativa Japonica Group cDNA clone:J023039P13, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015634503.1 | 0.0 | auxin response factor 3 isoform X2 | ||||
| Swissprot | Q5JMM1 | 0.0 | ARFC_ORYSJ; Auxin response factor 3 | ||||
| TrEMBL | A0A0D3EU34 | 0.0 | A0A0D3EU34_9ORYZ; Auxin response factor | ||||
| STRING | OBART01G31160.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5113 | 27 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33860.1 | 1e-148 | ARF family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



