 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB02G22950.1 | ||||||||
| Common Name | LOC102710275 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 401aa MW: 43480.1 Da PI: 6.6544 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.1 | 3.5e-32 | 234 | 289 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
kpr++W+peLH+rF++a++qLGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
OB02G22950.1 234 KPRRCWAPELHRRFLQALQQLGGSHVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 289
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.508 | 231 | 291 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.4E-17 | 232 | 292 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-28 | 232 | 292 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.3E-27 | 234 | 289 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.8E-6 | 236 | 287 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 401 aa Download sequence Send to blast |
MEVVDHAERD LARRRCREYL LALEEERRKI QVFQRELPLC FDLVTQTIEG MRSQMDGVGS 60 EETVSDHGPP PVLEEFIPLK PSLSLSSSEE ESNHAASDKA GKEEGAQTSE RHSSPQTLPE 120 AKRVTPDWLQ SVQLWSQQPQ QPSSPSQTPA KDLPCKPVAL NARKTGGAFQ PFEKEKRAEL 180 PASSTTAAAS STVVGDSGDK PADDTDKPRE TDNDGMDGKD KDKEKESQSQ PHRKPRRCWA 240 PELHRRFLQA LQQLGGSHVA TPKQIRELMK VDGLTNDEVK SHLQKYRLHT RRPSSTGQSS 300 AAAGVAAPPA PQFVVVGSIW VPPPEYAAAA AAQQQQVQLA AADASVSANP VYAPVAMLAP 360 GLQPHSHREQ RQQQGQRHSG SEGSGDTGGG RSSSPAVSSS S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 8e-13 | 234 | 287 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-13 | 234 | 287 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-13 | 234 | 287 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-13 | 234 | 287 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-12 | 234 | 287 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that may play a role in response to nitrogen. May be involved in a time-dependent signaling for transcriptional regulation of nitrate-responsive genes. Binds specifically to the DNA sequence motif 5'-GAATC-3' or 5'-GAATATTC-3'. Represses the activity of its own promoter trough binding to these motifs. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
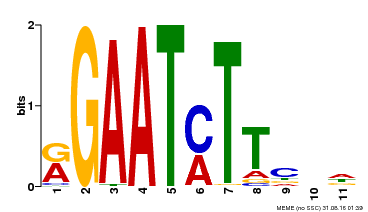
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB02G22950.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by nitrate. {ECO:0000269|PubMed:23324170}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK064355 | 0.0 | AK064355.1 Oryza sativa Japonica Group cDNA clone:002-108-B01, full insert sequence. | |||
| GenBank | AK101809 | 0.0 | AK101809.1 Oryza sativa Japonica Group cDNA clone:J033066M05, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015689147.1 | 0.0 | PREDICTED: probable transcription factor KAN2 | ||||
| Swissprot | Q6Z869 | 1e-179 | NIGT1_ORYSJ; Transcription factor NIGT1 | ||||
| TrEMBL | J3LCC8 | 0.0 | J3LCC8_ORYBR; Uncharacterized protein | ||||
| STRING | OB02G22950.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6470 | 37 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 4e-49 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 102710275 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



