 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OB01G18800.1 | ||||||||
| Common Name | LOC102707619 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 618aa MW: 67153.6 Da PI: 7.243 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.3 | 1.2e-12 | 464 | 510 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
OB01G18800.1 464 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITYITDLQ 510
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.9E-50 | 60 | 245 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.831 | 460 | 509 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.80E-15 | 463 | 514 | No hit | No description |
| SuperFamily | SSF47459 | 1.13E-18 | 463 | 526 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.7E-18 | 464 | 522 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.8E-10 | 464 | 510 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-16 | 466 | 515 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 618 aa Download sequence Send to blast |
MVMKMEAEED GANGGTGGTW TEEDRALSAS VLGTDAFAYL TKGGGAISEG LVAASLPVDL 60 QNRLQELVES ERPGVGWNYA IFWQLSRTKS GDLVLGWGDG SCREPRDGEV GAAASADNDE 120 AKQRMRKRVL QRLHSAFGGV DEEDYAPGID QVTDTEMFFL ASMYFAFPRR AGGPGQVFAA 180 GVPLWIPNTE RSVFPANYCY RGHLANAAGF RTIVLVPFET GVLELGSMQQ VAESSDTLQT 240 IRSVFAGAIG NKAGVQRHEG NGPADKSPGL AKIFGKDLNL GRPSAGPGAG VSKADERSWE 300 QRIAAGGSSL LPNVQRGLQN FTWSQARGLN SHQQKFGNGL LIVSNEATPR NNGVVESPTA 360 TQFQLQKAPP LQKLPKLQKP QQLVKPQQLV SQQQLQPQAP RQIDFSAGSS SKPGVLTKRP 420 AGIDGESAEV DGLCKDEGPP PAIEDRRPRK RGRKPANGRE EPLNHVEAER QRREKLNQRF 480 YALRAVVPNI SKMDKASLLG DAITYITDLQ KKLKEMEAER ERFLESGMVD PRERTPRPEV 540 DIQVVQDEVL VRVMSPMENH PVKTVFQAFE EAEVHAGESK ITSNNGTAVH SFIIKCPGAE 600 QQTREKVIAA VSRAMSSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 7e-28 | 458 | 520 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 445 | 453 | RRPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
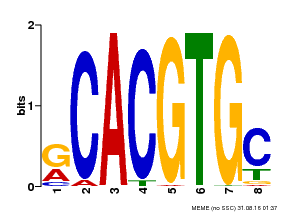
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | OB01G18800.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP012612 | 0.0 | CP012612.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 4 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006643964.1 | 0.0 | PREDICTED: transcription factor bHLH13 | ||||
| TrEMBL | J3KY24 | 0.0 | J3KY24_ORYBR; Uncharacterized protein | ||||
| STRING | OB01G18800.1 | 0.0 | (Oryza brachyantha) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 3e-71 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 102707619 |



