 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Neem_4530_f_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Meliaceae; Azadirachta
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 577aa MW: 64668.6 Da PI: 5.1242 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 170.2 | 6.5e-53 | 74 | 205 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp..kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevls 94
lp GfrF+Ptdeel+++yL+ k++g++ e+ evi+e+d++k+ePwdLp + +k+++ ew+fF++rd+ky++g r+nrat++gyWkatgkd+++ s
Neem_4530_f_1 74 LPLGFRFRPTDEELINHYLRLKINGRDSEV-EVIPEIDVCKWEPWDLPglSVIKTDDPEWFFFCPRDRKYPNGLRSNRATDAGYWKATGKDRTIRS 168
699************************999.99***************6447888899************************************** PP
NAM 95 k...kgelvglkktLvfykgrapkgektdWvmheyrl 128
+ +++ +g+kktLvfy+grapkge+t+W+mheyr+
Neem_4530_f_1 169 RksgSNNCIGMKKTLVFYRGRAPKGERTNWIMHEYRA 205
96655566***************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 7.32E-62 | 63 | 228 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.756 | 74 | 228 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.2E-27 | 75 | 204 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0071470 | Biological Process | cellular response to osmotic stress | ||||
| GO:1900426 | Biological Process | positive regulation of defense response to bacterium | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005789 | Cellular Component | endoplasmic reticulum membrane | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 577 aa Download sequence Send to blast |
MSRDQSVLRN GIPAQERRSK AGESRALCFK QLKRPFITKR DSTRRGVQKA TRFGHEMLTC 60 RKCNTMTVLS VESLPLGFRF RPTDEELINH YLRLKINGRD SEVEVIPEID VCKWEPWDLP 120 GLSVIKTDDP EWFFFCPRDR KYPNGLRSNR ATDAGYWKAT GKDRTIRSRK SGSNNCIGMK 180 KTLVFYRGRA PKGERTNWIM HEYRATDKDL QVTKPGQAAF VLCRLFRKPE EKNDVEAQQT 240 ESPDMTKSSP DDTSSDLVQE TPISDYECPG VGENPYQYEL PSNGEIDCKI FSPMQSNPDL 300 GSLDGTNEQD VSLSDLLDEV FNNHDEFSCE ESNGQKELAI ETKMQLPGEI YMSQTISSEN 360 FHVKDNSTFS VTDTGVAQLQ HRDLAMGSSG WFNDHLESKG SFGGYGDNSG TQLSSSNNSA 420 MGSFCGNFNN YDLSTTLKNP VDNSGDFAGG TGIKIRTRQP QHRPNSFNSV TQGTAPGRLR 480 LQTELSSAPV GTGEQKDVSL SKEEDEVQSV VTEIDEPAEQ SQTSDGESQV LELDETKELT 540 QEASTKLRLR VKRDGEIWHS QDGAISVKRG TDCTSWK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 1e-45 | 69 | 228 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 1e-45 | 69 | 228 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 1e-45 | 69 | 228 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 1e-45 | 69 | 228 | 12 | 165 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 1e-45 | 69 | 228 | 12 | 165 | NAC domain-containing protein 19 |
| 4dul_B | 1e-45 | 69 | 228 | 12 | 165 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP) (PubMed:18443413, PubMed:24329768). Calmodulin-regulated transcriptional repressor. Binds several synthetic promoters with randomly selected binding sites (PubMed:17947243). Functions synergistically with SNI1 as negative regulator of pathogen-induced PR1 expression and basal resistance to a virulent strain of P.syringae. Binds directly to the promoter of the PR1 gene (PubMed:22826500). Acts as positive regulator of innate immunity. Involved in the effector-triggered immunity (ETI) induction of immunity-related gene expression (PubMed:24329768). Mediates osmotic stress signaling in leaf senescence by up-regulating senescence-associated genes (PubMed:18443413). {ECO:0000269|PubMed:17947243, ECO:0000269|PubMed:18443413, ECO:0000269|PubMed:22826500, ECO:0000269|PubMed:24329768}. | |||||
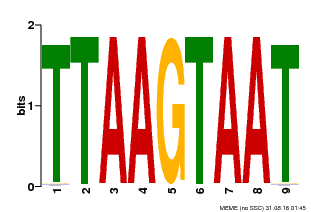
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00108 | PBM | Transfer from AT4G35580 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006439176.1 | 0.0 | protein NTM1-like 9 | ||||
| Swissprot | F4JN35 | 1e-128 | NTL9_ARATH; Protein NTM1-like 9 | ||||
| TrEMBL | V4TR99 | 0.0 | V4TR99_9ROSI; Uncharacterized protein | ||||
| STRING | XP_006439176.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5594 | 28 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G35580.2 | 1e-130 | NAC transcription factor-like 9 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



