 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.I01197.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 355aa MW: 39776.3 Da PI: 9.7832 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 48.8 | 1.9e-15 | 46 | 133 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkm.rergferspk...qCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
+W++ + +Li+a e+ +l+rg+lk+++W++v++ + +++ + ++pk qCk++++ ++k+yk +k+ ++ + s++++fd+l+
Migut.I01197.1.p 46 CWSEAATAVLIDAWGERYLELSRGNLKQKHWKDVADIVcSREDYAKTPKtdiQCKNRIDTVKKKYKTEKAKIASG--GGPSKWRFFDKLD 133
7**********************************976677888777754449********************97..67778******97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF13837 | 2.2E-25 | 44 | 134 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009793 | Biological Process | embryo development ending in seed dormancy | ||||
| GO:0010431 | Biological Process | seed maturation | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 355 aa Download sequence Send to blast |
MEEDDEIQSP TPNGRITVTV ASAPPSNSLT LALPIQPQKT PGREDCWSEA ATAVLIDAWG 60 ERYLELSRGN LKQKHWKDVA DIVCSREDYA KTPKTDIQCK NRIDTVKKKY KTEKAKIASG 120 GGPSKWRFFD KLDVLIGPTP SKSNSTPPPP MDLYQRGVPM GIPVGIRSSS SVLKRPSFDS 180 EDESEPENSA DSSDGLPPDT YNNNNKRGKI QKMEMPTVNN ANGSSSQRRD GGGMRGNGRK 240 NKEESNWANS MKELTRAVLK FGEAYEQAET AKLQQLVEME KQRMKFAKEL ELQRMQYFMK 300 TQMELSQIKN RSSANTNNHH HHHHRRVISN NNNNDANNNN TKKNNNTKNS DSSN* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription regulator that may repress the maturation program during early embryogenesis. {ECO:0000269|PubMed:21330492}. | |||||
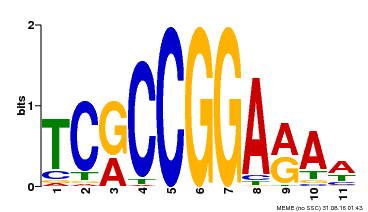
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00356 | DAP | Transfer from AT3G14180 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.I01197.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012856279.1 | 0.0 | PREDICTED: trihelix transcription factor ASIL2-like | ||||
| Swissprot | Q9LJG8 | 7e-66 | ASIL2_ARATH; Trihelix transcription factor ASIL2 | ||||
| TrEMBL | A0A2G9I2A2 | 1e-146 | A0A2G9I2A2_9LAMI; Transcription factor GT-2 | ||||
| STRING | Migut.I01197.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10332 | 18 | 23 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G14180.1 | 3e-57 | sequence-specific DNA binding transcription factors | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.I01197.1.p |



