 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.H00617.1.p | ||||||||
| Common Name | LOC105971404, MIMGU_mgv1a011591mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 278aa MW: 31066 Da PI: 6.7234 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 99.8 | 5e-31 | 50 | 164 | 2 | 89 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt..............ssssaseceaessss 80
g+kdrhsk++T +g+RdRRvRls+++a++++dLqd+LG+ ++sk ++WLl ak +i+el+ + + ++s++s
Migut.H00617.1.p 50 FGGKDRHSKVCTVRGLRDRRVRLSVPTAIQLYDLQDRLGLNQPSKVVDWLLDIAKNEIDELPPLqipqgmminwnssnV------VPEHSRRS 136
689*************************************************************555555543222220......22222222 PP
TCP 81 sasnsssg.........................k 89
+ + +
Migut.H00617.1.p 137 E------DinlfggnnnklkevdtsnnrdeaetN 164
2......022333332222222222221222220 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.8E-29 | 52 | 161 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 30.555 | 52 | 110 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0031347 | Biological Process | regulation of defense response | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| GO:0009507 | Cellular Component | chloroplast | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 278 aa Download sequence Send to blast |
MSSTSNNTDN NPAAKNLFKA ATTSTTTTAA SSWSNTRLKD PRIVRVSRAF GGKDRHSKVC 60 TVRGLRDRRV RLSVPTAIQL YDLQDRLGLN QPSKVVDWLL DIAKNEIDEL PPLQIPQGMM 120 INWNSSNVVP EHSRRSEDIN LFGGNNNKLK EVDTSNNRDE AETNWTRSSD EGALYNNVSA 180 YNSYVRWDNY PSNMLTLSHD QQHNNHNPTK GHALITSAHN QLNDDWHNFS LQLPSYFATT 240 GAEFDAEKAN YDQRSQLESS SVLCTANLLP SQNNDGS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 1e-18 | 57 | 111 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 1e-18 | 57 | 111 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164). Binds to the 3'-ACC-5' repeats in the light-responsive promoter (LRP) of psbD, and activates its transcription. Participates in ovule develpment (PubMed:25378179). {ECO:0000269|PubMed:11161017, ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:25378179}. | |||||
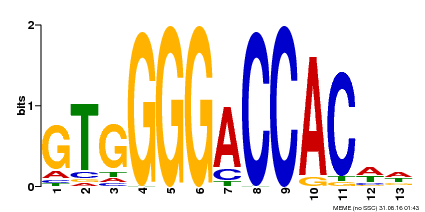
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00326 | DAP | Transfer from AT3G02150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.H00617.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012851715.1 | 0.0 | PREDICTED: transcription factor TCP13-like | ||||
| Swissprot | Q9S7W5 | 1e-44 | TCP13_ARATH; Transcription factor TCP13 | ||||
| TrEMBL | A0A022RZ06 | 0.0 | A0A022RZ06_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.H00617.1.p | 0.0 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA821 | 24 | 99 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G02150.1 | 4e-47 | plastid transcription factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.H00617.1.p |
| Entrez Gene | 105971404 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



