 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.D01635.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 275aa MW: 30803.2 Da PI: 8.8248 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 95.1 | 1.3e-29 | 54 | 163 | 2 | 89 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt..................ssssaseceae 76
g+kdrhsk++T +g+RdRR+Rls+++a++++dLqd+LG+ ++sk ++WL+++ i++l+ + ++ssa
Migut.D01635.2.p 54 FGGKDRHSKVCTVRGLRDRRIRLSVPTAIQLYDLQDRLGLSQPSKVVDWLIEATMNDIDKLPPLpmlptnfpqfhhqpitidNQSSA------ 140
689*************************************************************66666555443333322211111...... PP
TCP 77 sssssasnsssg..............k 89
+++ n+s + +
Migut.D01635.2.p 141 ---NNTTNQS-SlshflasnypfarlD 163
...1111111.1234443322232221 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51369 | 28.31 | 56 | 114 | IPR017887 | Transcription factor TCP subgroup |
| Pfam | PF03634 | 3.2E-27 | 56 | 168 | IPR005333 | Transcription factor, TCP |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 275 aa Download sequence Send to blast |
MTMNPKEKDF LAKKELCGDG NNKKIIATST IPSSSRQWSS THRNPRIVRV SRAFGGKDRH 60 SKVCTVRGLR DRRIRLSVPT AIQLYDLQDR LGLSQPSKVV DWLIEATMND IDKLPPLPML 120 PTNFPQFHHQ PITIDNQSSA NNTTNQSSLS HFLASNYPFA RLDKERDQEN QENDIQEGFG 180 GIFAQNLFPL TNINSQSSFA NLPFNNAYNF HWDPASLSLS QFGGGYSFTT NQTAVDEENN 240 NNNLSSNNSQ LYFCPSASNI VPSVIPNNSR THRT* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 7e-20 | 61 | 115 | 1 | 55 | Putative transcription factor PCF6 |
| 5zkt_B | 7e-20 | 61 | 115 | 1 | 55 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Plays a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164). Participates in ovule develpment (PubMed:25378179). {ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:25378179}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
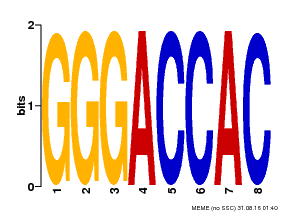
| Motif ID | Method | Source | Motif file |
| MP00008 | PBM | Transfer from 496250 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.D01635.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012854041.1 | 0.0 | PREDICTED: transcription factor TCP5 | ||||
| Swissprot | Q9FME3 | 2e-45 | TCP5_ARATH; Transcription factor TCP5 | ||||
| TrEMBL | A0A022Q7L4 | 1e-116 | A0A022Q7L4_ERYGU; Uncharacterized protein | ||||
| STRING | Migut.D01635.1.p | 0.0 | (Erythranthe guttata) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G60970.1 | 9e-47 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.D01635.2.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



