 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly0502s0001.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 565aa MW: 58206.6 Da PI: 8.3142 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36.2 | 1.1e-11 | 506 | 551 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR+ri +++ L+el+P++ K +Ka++L + +eY+k Lq
Mapoly0502s0001.1.p 506 HSIAERLRRERIAERMKALQELVPNS-----NKTDKASMLDEIIEYVKFLQ 551
99***********************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 4.45E-16 | 499 | 557 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.352 | 501 | 550 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.0E-15 | 503 | 555 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 7.83E-13 | 505 | 555 | No hit | No description |
| Pfam | PF00010 | 4.8E-9 | 506 | 551 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 8.5E-14 | 507 | 556 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048767 | Biological Process | root hair elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 565 aa Download sequence Send to blast |
MAAQQGVVVG VSSSLSSAHR LLLGGGPGGP SSPGPTQSPS SSCCPTSCSP SLQASARTLL 60 GCSLPPAAAA AAAAAAGSAA LASAAALVVS TQTCSSSQAQ QYQYQQRRCN PAAAAFHHHQ 120 LQQQQAQPPF LSPPALQALL PSGAHSEMQP ASMLGSRGPV PGVSSARSSL SNAALAMPSL 180 LSPPHQQPAL QFEQAEQTPP MDEFLEQMFS LPSWSEMAGA GRAPWEFNAA AAGAATRLGV 240 DAQKLFTMSL LPPGLLGGPH TSTTTTNNNN TNNNNATSNN NNNHSGRDLQ DGADSAHVGD 300 SVLGYSNDDH LLGARLRGLQ ETPSSVPGIG TRAGATLTRS PSVGSSGSEG SGGPQSPPPI 360 APTWRQPYVG GVPPLPLGLG QAKPETLLRG DSGEAHLLGK RPRDEDEVSM REAHTGNNVR 420 DAPQGLFASF TGGQVTQNQV PRSAGQQSHQ QGNGASSQGV QGQGQALPTY GSPSITATTQ 480 PHSNSGVAVA ARPRVRARRG QATDPHSIAE RLRRERIAER MKALQELVPN SNKTDKASML 540 DEIIEYVKFL QLQVKVSCTG SLSLS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 510 | 515 | RLRRER |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
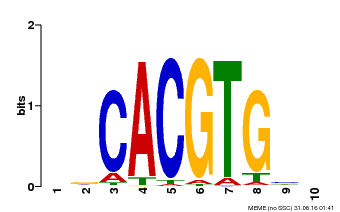
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A2R6VYU3 | 0.0 | A0A2R6VYU3_MARPO; Uncharacterized protein (Fragment) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP58 | 16 | 313 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58010.1 | 1e-19 | LJRHL1-like 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly0502s0001.1.p |



