 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.17G060600.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 405aa MW: 46635.9 Da PI: 4.7088 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 112.6 | 2.6e-35 | 14 | 105 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkk 91
Fl+k+ye+++d+++++++sws++++sf+v+++ efa+++Lp++Fkh+nf+SF+RQLn+YgF+k++ e+ weF+++ F +g+
Manes.17G060600.1.p 14 FLSKTYEMVDDPSTDSVVSWSQSNKSFIVWNPPEFARDLLPRFFKHNNFSSFIRQLNTYGFRKIDPEQ---------WEFANEDFIRGQP 94
9***************************************************************9998.........************* PP
XXXXXXXXXXX CS
HSF_DNA-bind 92 ellekikrkks 102
+l+++i+r+k
Manes.17G060600.1.p 95 HLMKNIHRRKP 105
********985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46785 | 4.9E-36 | 9 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 3.0E-64 | 10 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene3D | G3DSA:1.10.10.10 | 6.0E-39 | 10 | 103 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 1.7E-31 | 14 | 103 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 3.0E-20 | 14 | 37 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 3.0E-20 | 52 | 64 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 53 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 3.0E-20 | 65 | 77 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 405 aa Download sequence Send to blast |
MDESQGSSNS LPPFLSKTYE MVDDPSTDSV VSWSQSNKSF IVWNPPEFAR DLLPRFFKHN 60 NFSSFIRQLN TYGFRKIDPE QWEFANEDFI RGQPHLMKNI HRRKPVHSHS MQNLQGQGSY 120 TLTDPERQTL RDDIERLQRE KEVLISDLQR HEQERKGFEM QMQALKEKLQ QMERRQQTMV 180 SFVAQVLQKP GLALNLMSQL EPGHDRKRRL PRIGYFYDEA SMEENQMVTC QTRENVDSNT 240 AALSNMEQFE QLESSLTFWE SIANDVENKM QSNSNLELDE STSCAESPAI SCVQFSVDIR 300 PKSPAIDMNS EPAVASVPEP EPLKEQTTGT APTVAAGVND IFWEQFLTEN PGSTDAQEVQ 360 SERKDSGERK NEMKPSDQGK FWWNMRNVNN LAEQMEHLTP AERT* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 6e-25 | 4 | 103 | 20 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 6e-25 | 4 | 103 | 20 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 6e-25 | 4 | 103 | 20 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 204 | 209 | DRKRRL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds DNA sequence 5'-AGAAnnTTCT-3' known as heat shock promoter elements (HSE). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
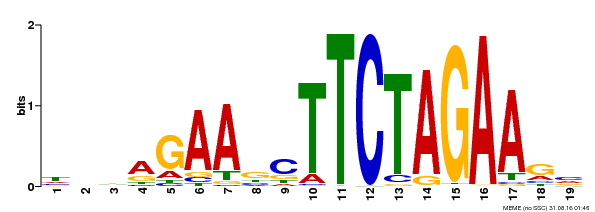
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.17G060600.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021599108.1 | 0.0 | heat stress transcription factor A-4a isoform X1 | ||||
| Refseq | XP_021599109.1 | 0.0 | heat stress transcription factor A-4a isoform X1 | ||||
| Swissprot | O49403 | 1e-145 | HFA4A_ARATH; Heat stress transcription factor A-4a | ||||
| TrEMBL | A0A2C9U5F3 | 0.0 | A0A2C9U5F3_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_008941m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3568 | 33 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 1e-143 | heat shock transcription factor A4A | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.17G060600.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



