 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.08G069300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 842aa MW: 96661.3 Da PI: 6.9823 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.7 | 4.1e-28 | 79 | 166 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHela 90
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e++ + ++r++++++t+Cka+++vk++ dgkw ++++ +eHnHel
Manes.08G069300.1.p 79 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGITPESDGS---NSRRSSVKKTDCKASMHVKRRPDGKWIIHEFIKEHNHELL 165
5****************************************999887...889999********************************97 PP
FAR1 91 p 91
p
Manes.08G069300.1.p 166 P 166
5 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.0E-25 | 79 | 166 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 6.1E-25 | 287 | 379 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.322 | 566 | 602 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.1E-4 | 575 | 600 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 9.5E-8 | 577 | 604 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 842 aa Download sequence Send to blast |
MIIMTGRNQH AVHDMVSLQE NVLVGDARGE NMVDIVDEAQ NGDSGVVDFP KRAVTLFEED 60 TDFEPCNGIE FESHEAAYSF YQEYAKSMGF TTSIKNSRRS KKSKEFIDAK FACSRYGITP 120 ESDGSNSRRS SVKKTDCKAS MHVKRRPDGK WIIHEFIKEH NHELLPALAY HFRIHRNVKL 180 AEKNNIDILH AVSERTRKVY VEMLRQSGGF KNVSLVQNDN GTQFEKGRPL ALEEGDAQAL 240 LEYFKHIKKE NPNFFYAIDL NEEQRLRNLF WVDAKSRNDY ISFNDAVSFD TFYAKYHEKL 300 PFAPFVGVNH HCQLILLGCA LVADDSMSTF VWLLRTWLRA MGGKAPKVII TDLDNTLKAA 360 IEEVFPNTRH CFSLWHIFEK MPETLSHVIK RHGDFMPEFH KCVFKSWTDE EFDMRWWTMV 420 TQFELQDNEW IQSLYDDRKK WVPTYMEDTF LAGIAASQRS ESVNSFFDKY IHRKITLKEF 480 MKQYGLILQN RYEEEAIADF DTSHKQPALK SPSPWEKQMS MVYTHAIFKK FQVEVLGVVG 540 CHPKKEREDG TNVTFRVQDC EKNEYFLVSW NQSKSAVSCL CRLFEYKGFL CRHALIVLQI 600 CGLSSIPTHY ILKRWTKDAK SGQQLAEGKA RIETRVQRYN ALCKLAIEMS EEGSLCEQSY 660 NIAFRTLVET LKNCVNVNNR NNSALEPGSQ SFAPCEAEDE NQGNFSKSSK KKNPIRKRKV 720 QADPDIMLVE AQESLQQMEN LGSDTLNGYY GAQQNAQGLV QLNLMEPPHD GYYVSQPSIQ 780 GLGQLNSIAP GHDGFFGTQQ SINGMGQFDY RPPTSFNYSL QEDSHLRSSQ LHGSAPRHTQ 840 G* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
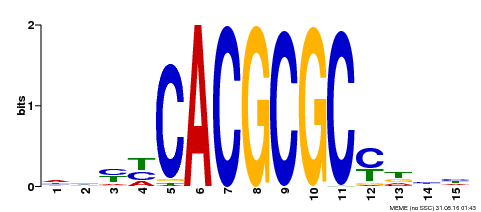
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.08G069300.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021620401.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A2C9VE37 | 0.0 | A0A2C9VE37_MANES; Uncharacterized protein | ||||
| STRING | POPTR_0006s02140.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.08G069300.1.p |



