|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
MLOC_66348.1 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
| Family |
NAC |
| Protein Properties |
Length: 337aa MW: 37691.2 Da PI: 5.7209 |
| Description |
NAC family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| MLOC_66348.1 | genome | IBSC | View CDS |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | NAM | 161.2 | 3.9e-50 | 38 | 169 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk.. 95
pG+rFhPtdeelv++yL++kv++k+l++ evi+e+diyk++PwdLp +++ ++ekewyfF+ r +ky+++ r+nr+t sg+Wkatg d++++s+
MLOC_66348.1 38 LPGYRFHPTDEELVTFYLRRKVARKSLRI-EVIREIDIYKHDPWDLPkASTVGGEKEWYFFCLRGRKYRNSIRPNRVTGSGFWKATGIDRPIYSAaa 133
69***************************.99***************445556889**************************************988 PP
NAM 96 ...kgelvglkktLvfykgrapkgektdWvmheyrl 128
+g +glkk+Lv+y+g+a kg+ktdW+mhe+rl
MLOC_66348.1 134 graSGVSIGLKKSLVYYRGSAGKGTKTDWMMHEFRL 169
7777777***************************98 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0005992 | Biological Process | trehalose biosynthetic process |
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated |
| GO:0006561 | Biological Process | proline biosynthetic process |
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process |
| GO:0010120 | Biological Process | camalexin biosynthetic process |
| GO:0042538 | Biological Process | hyperosmotic salinity response |
| GO:1900056 | Biological Process | negative regulation of leaf senescence |
| GO:0003677 | Molecular Function | DNA binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 337 aa
Download sequence Send
to blast |
MVAKADMTVM EMEKATSDGE VVLGGAGDDD DEEEDVVLPG YRFHPTDEEL VTFYLRRKVA 60
RKSLRIEVIR EIDIYKHDPW DLPKASTVGG EKEWYFFCLR GRKYRNSIRP NRVTGSGFWK 120
ATGIDRPIYS AAAGRASGVS IGLKKSLVYY RGSAGKGTKT DWMMHEFRLP PAAAAAATNS 180
SPSMQEAEVW TICRIFRRTI TYRKQQQQPW RPAPAVSAAD SSSNTGSFES SEVSDEYINC 240
LQAPAAAAPC IPEQHQYVSQ TGTVDSGNYF YMDTMHNQQQ VQGQWNAPPA APVPEHKPQS 300
QLSAANDFYK VEGYLEEIAR MMEVTDPAGF YDCRSYA
|
| Functional Description ? help
Back to Top |
| Source |
Description |
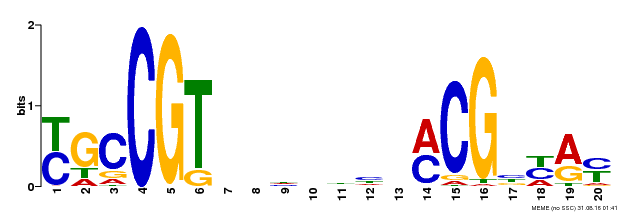
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | AK374728 | 0.0 | AK374728.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3074N05. |
| Publications
? help Back to Top |
- Shahnejat-Bushehri S,Nobmann B,Devi Allu A,Balazadeh S
JUB1 suppresses Pseudomonas syringae-induced defense responses through accumulation of DELLA proteins.
Plant Signal Behav, 2016. 11(6): p. e1181245
[PMID:27159137] - Shahnejat-Bushehri S,Tarkowska D,Sakuraba Y,Balazadeh S
Arabidopsis NAC transcription factor JUB1 regulates GA/BR metabolism and signalling.
Nat Plants, 2016. 2: p. 16013
[PMID:27249348] - Shahnejat-Bushehri S, et al.
Arabidopsis NAC Transcription Factor JUNGBRUNNEN1 Exerts Conserved Control Over Gibberellin and Brassinosteroid Metabolism and Signaling Genes in Tomato.
Front Plant Sci, 2017. 8: p. 214
[PMID:28326087] - Sakuraba Y,Bülbül S,Piao W,Choi G,Paek NC
Arabidopsis EARLY FLOWERING3 increases salt tolerance by suppressing salt stress response pathways.
Plant J., 2017. 92(6): p. 1106-1120
[PMID:29032592] - Ebrahimian-Motlagh S, et al.
JUNGBRUNNEN1 Confers Drought Tolerance Downstream of the HD-Zip I Transcription Factor AtHB13.
Front Plant Sci, 2017. 8: p. 2118
[PMID:29326734]
|