 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_57622.4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 358aa MW: 38626.2 Da PI: 6.6269 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 26.5 | 1.2e-08 | 170 | 217 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR+ri +++ L++l+P + +k Ka +L + ++Y++sLq
MLOC_57622.4 170 SHSLAERVRRERISQRMKFLQDLVPGC----NKVIGKALMLDEIINYVQSLQ 217
8*************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 3.14E-16 | 164 | 231 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 8.10E-10 | 164 | 221 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.6E-15 | 166 | 229 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 14.964 | 166 | 216 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 7.2E-6 | 170 | 217 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.3E-8 | 172 | 222 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 358 aa Download sequence Send to blast |
MEGRAGGGSG DYISSLLTSS PMLDFGMLDG AVTAGGDCLD KFCGDPGFAE RAARLSSFNG 60 QHLAPGLLGM LPPAPGGEFG GSREASSVSD PASAMKDANA KKRKAPASKG KAKEPSLSTS 120 CQVGEHKEPD AKRCRTGDAE KKGKGKNAKP VEPPKDYVHV RARRGQATDS HSLAERVRRE 180 RISQRMKFLQ DLVPGCNKVI GKALMLDEII NYVQSLQRQV EFLSMKLATV NPLDFSNLPT 240 LLHKDMYGPS ASSVFSLESS SSAFPFSDQG DVFQSFLPNS MESQCTLNQL DLALSQATNA 300 AQYGFQDATA STNLQQRNFW EDDLQSVFHV DNRQSQDNGV SAESFHGDLQ AGQMKMEF |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00036 | PBM | Transfer from AT5G48560 | Download |
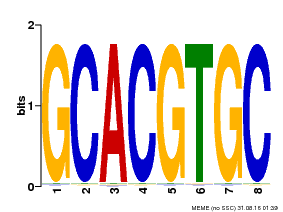
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_57622.4 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK367580 | 0.0 | AK367580.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2059B18. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020146738.1 | 0.0 | transcription factor bHLH78-like isoform X2 | ||||
| TrEMBL | M0X7F0 | 0.0 | M0X7F0_HORVV; Uncharacterized protein | ||||
| STRING | MLOC_57622.3 | 0.0 | (Hordeum vulgare) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G07340.1 | 8e-60 | bHLH family protein | ||||



