 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C025844P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 693aa MW: 75312.9 Da PI: 6.6596 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 21.3 | 4.8e-07 | 276 | 296 | 5 | 26 |
--SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHg 26
+C+ f++ GtC+ Gd C +aHg
MELO3C025844P1 276 PCPEFRK-GTCRQGDACEYAHG 296
8******.*************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF48403 | 3.58E-13 | 72 | 150 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.175 | 75 | 150 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 8.6E-7 | 75 | 142 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0049 | 75 | 105 | IPR002110 | Ankyrin repeat |
| Gene3D | G3DSA:1.25.40.20 | 1.1E-14 | 76 | 168 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.75E-12 | 77 | 150 | No hit | No description |
| PROSITE profile | PS50088 | 11.648 | 110 | 145 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.23 | 110 | 142 | IPR002110 | Ankyrin repeat |
| SMART | SM00356 | 5.4E-5 | 271 | 297 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 12.784 | 276 | 298 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.2E-4 | 276 | 296 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 8.9E-4 | 304 | 334 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 30 | 306 | 329 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 7.103 | 306 | 330 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 693 aa Download sequence Send to blast |
MMCNGCNRKS ASPNFAMDGG FKRGGLGFCI SVLLEFSATD DLIGFRSAVE EDGHDIDEAS 60 LWYGRIFGSK KMGYEERTPL MVAAMFGSLN VLSYILHSGR VDVNRACGSD GVTTLHCAVA 120 GGSAVVDQVV KLLLDASADV SAVDANGNRP GDLIAPDFTS AFHSRKKTLQ QLLNGHEEGL 180 SSSEAIFYER ETLEPLELST LRASRDGTEK KEYPVDLSLP DIKNGIYSTD EFRMYTFKIK 240 PCTRAYSHDW TECPFVHPGE NARRRDPRKY HYSCVPCPEF RKGTCRQGDA CEYAHGIFEC 300 WLHPAQYRTR LCKDETGCTR KVCFFAHKPE ELRPLYASTG SAVLSPRSIC GSSLDIASIS 360 PLTLGSPSAL IPPSSTPPLT PSGVSSPMGG TVWQTQCNIA PPTLHLPGSR LKASLSARDV 420 DLDVELLGLE SQRRRQQQLM DEMSCLPSPS RWNNGIPTPA SFPSPRSRNG ELNGLGGMKP 480 TKLEDIFGSV DPAILPQLQG LSLDSVGSQV QSPSGIQMRQ SLNQSFLSSY GNSIGSPPPR 540 LSQPSVSTAA SVLSSRAAAF AKRSQSFIER SMVSRHTGLS PPATSTTTMP LNLSDWGSPD 600 GKLDWGIRGE ELNKLRKSAS FGFRNNCTSS PVTSTMHTTA PEPDVSWVQS LVKDAPSENA 660 VQLSMDEQQQ LLCHINNGDS KQMYMEQEQL VA* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00566 | DAP | Transfer from AT5G58620 | Download |
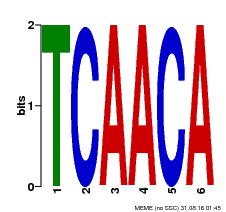
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681919 | 0.0 | LN681919.1 Cucumis melo genomic scaffold, anchoredscaffold00084. | |||
| GenBank | LN713265 | 0.0 | LN713265.1 Cucumis melo genomic chromosome, chr_11. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008464034.1 | 0.0 | PREDICTED: zinc finger CCCH domain-containing protein 66 | ||||
| Swissprot | Q9LUZ4 | 0.0 | C3H66_ARATH; Zinc finger CCCH domain-containing protein 66 | ||||
| TrEMBL | A0A1S3CL13 | 0.0 | A0A1S3CL13_CUCME; zinc finger CCCH domain-containing protein 66 | ||||
| STRING | XP_008464034.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1796 | 34 | 90 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58620.1 | 0.0 | C3H family protein | ||||



