 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C022694P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 371aa MW: 40957 Da PI: 4.9899 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.4 | 9.7e-20 | 115 | 164 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+G+r+++ +g+W+AeIrdp++ +r++lg+f taeeAa+a++a +++++g
MELO3C022694P1 115 QYRGIRQRP-WGKWAAEIRDPRK---GARVWLGTFNTAEEAARAYDAEARRIRG 164
69*******.**********954...4*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 3.49E-33 | 115 | 173 | No hit | No description |
| PROSITE profile | PS51032 | 24.026 | 115 | 172 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.5E-32 | 115 | 173 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.18E-22 | 115 | 173 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 2.1E-37 | 115 | 178 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 8.5E-12 | 116 | 127 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.4E-12 | 116 | 164 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 8.5E-12 | 138 | 154 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0070483 | Biological Process | detection of hypoxia | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 371 aa Download sequence Send to blast |
MCGGAIISDF IPPSCSNRVT ADHLWPNLKK PKSGKHSSPR SLRSRIFDVD DFEADFRDFK 60 DESDVEDEDD FSDIKPFLFS TPNSACSSTR GSSATKSLEF NEQAEKSANT KRKNQYRGIR 120 QRPWGKWAAE IRDPRKGARV WLGTFNTAEE AARAYDAEAR RIRGNKARVN FPDEPLPNTQ 180 KRKNSQKAKQ HIKENVKANQ HPNQNYSGTT GFLEVKPPTD QVGYMDSFPA SMDSSPSDDM 240 VMYFNSDEGS NSISCSGFGM GDHGVKTPEI SSVFSATDSE FTEDMHTRKK QRCSSGDAIT 300 AEDVGASAKT LSEELSAFES QMKLFQMPYL DGNWDDSMDA FLGGGATQDG GNSLDLWSFD 360 DLPAMAGGVF * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 5e-22 | 115 | 172 | 2 | 60 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in carotenoid biosynthesis regulation. Binds to the 5'-ATCTA-3' element present in the promoter of phytoene synthase (PSY) and phytoene desaturase (PDS). Involved in ethylene response and resistance to necrotrophic pathogens. Acts as a downstream regulator in the ethylene signaling pathway. Partially redundant with RAP2-12. {ECO:0000269|PubMed:17873090, ECO:0000269|PubMed:20357136, ECO:0000269|PubMed:22530619}. | |||||
| UniProt | Transcription factor involved in the activation of hypoxic gene expression and in ethylene response. Partially redundant with RAP2-2. Acts as a downstream regulator in the ethylene signaling pathway. {ECO:0000269|PubMed:22020282, ECO:0000269|PubMed:22530619}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
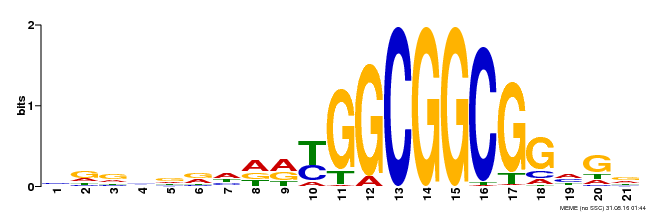
| Motif ID | Method | Source | Motif file |
| MP00204 | DAP | Transfer from AT1G53910 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by darkness, ethylene and Botrytis cinerea. {ECO:0000269|PubMed:20357136, ECO:0000269|PubMed:22530619}. | |||||
| UniProt | INDUCTION: Up-regulated by hypoxia but not by ethylene. {ECO:0000269|PubMed:22020282}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681829 | 0.0 | LN681829.1 Cucumis melo genomic scaffold, anchoredscaffold00054. | |||
| GenBank | LN713258 | 0.0 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008460339.1 | 0.0 | PREDICTED: ethylene-responsive transcription factor RAP2-12 | ||||
| Refseq | XP_008460340.1 | 0.0 | PREDICTED: ethylene-responsive transcription factor RAP2-12 | ||||
| Swissprot | Q9LUM4 | 1e-68 | RAP22_ARATH; Ethylene-responsive transcription factor RAP2-2 | ||||
| Swissprot | Q9SSA8 | 7e-69 | RA212_ARATH; Ethylene-responsive transcription factor RAP2-12 | ||||
| TrEMBL | A0A1S3CCQ5 | 0.0 | A0A1S3CCQ5_CUCME; ethylene-responsive transcription factor RAP2-12 | ||||
| STRING | XP_008460339.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4808 | 32 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G53910.3 | 3e-66 | related to AP2 12 | ||||



