 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C006700P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 125aa MW: 14594.8 Da PI: 9.8851 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.1 | 3.5e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rgrWT+eEde+l ++++ G g+W++ ++ g+ R++k+c++rw +yl
MELO3C006700P1 14 RGRWTAEEDEILTNYIQANGEGSWRSLPKNAGLLRCGKSCRLRWINYL 61
8*******************************99************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 21.098 | 9 | 65 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-19 | 12 | 59 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.7E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.4E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 3.47E-20 | 15 | 85 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.19E-8 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 5.6E-6 | 60 | 85 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50090 | 4.252 | 62 | 101 | IPR017877 | Myb-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 125 aa Download sequence Send to blast |
MVRAPCCEKV GLKRGRWTAE EDEILTNYIQ ANGEGSWRSL PKNAGLLRCG KSCRLRWINY 60 LRTDLKRGNI TTEEEQMIVK LHNVFWQQMV FNSGTFTWPD RQRNKKLLEL SSKPKNLHYD 120 TTFC* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Flavonol-specific transcription activator involved in the regulation of several genes of flavonoid biosynthesis. Activates the expression of CHS, CHI, F3H and FLS1. Controls flavonol biosynthesis mainly in the root (PubMed:17419845, PubMed:20731781). Confers tolerance to UV-B (PubMed:19895401). {ECO:0000269|PubMed:15923334, ECO:0000269|PubMed:17419845, ECO:0000269|PubMed:19895401, ECO:0000269|PubMed:20731781}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00668 | SELEX | Transfer from GRMZM2G016020 | Download |
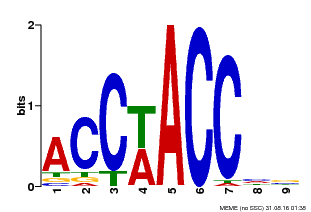
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By nitrogen deficiency, sucrose and UV LIGHT (PubMed:17053893, PubMed:9839469). Triggered by HY5 in response to light and UV-B (PubMed:19895401). {ECO:0000269|PubMed:17053893, ECO:0000269|PubMed:19895401, ECO:0000269|PubMed:9839469}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681848 | 4e-68 | LN681848.1 Cucumis melo genomic scaffold, anchoredscaffold00006. | |||
| GenBank | LN713260 | 4e-68 | LN713260.1 Cucumis melo genomic chromosome, chr_6. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004134499.1 | 1e-55 | PREDICTED: myb-related protein 308 | ||||
| Swissprot | O22264 | 2e-47 | MYB12_ARATH; Transcription factor MYB12 | ||||
| TrEMBL | A0A0A0L945 | 8e-56 | A0A0A0L945_CUCSA; Uncharacterized protein | ||||
| TrEMBL | A0A1S3AX58 | 6e-54 | A0A1S3AX58_CUCME; LOW QUALITY PROTEIN: transcription factor MYB12 | ||||
| STRING | XP_004134499.1 | 4e-55 | (Cucumis sativus) | ||||
| STRING | XP_004162204.1 | 4e-55 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47460.1 | 9e-50 | myb domain protein 12 | ||||



