 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C005072P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 296aa MW: 31212.6 Da PI: 6.7995 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 33.4 | 1e-10 | 12 | 55 | 4 | 45 |
S-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwq 45
W+ e d+ + +a + ++ + +W++Ia ++ g+t +++k+++
MELO3C005072P1 12 WSLEQDKAFENALASHPEDdsdRWEKIAVDVP-GKTVEEIKHHYE 55
********************************.**********96 PP
| |||||||
| 2 | Myb_DNA-binding | 42 | 2.2e-13 | 122 | 166 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+ l++ + ++G+g+W++I+r + +Rt+ q+ s+ qky
MELO3C005072P1 122 AWTEDEHRLFLLGLDKYGKGDWRSISRNFVVTRTPTQVASHAQKY 166
6*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 7.339 | 7 | 62 | IPR017884 | SANT domain |
| SMART | SM00717 | 5.1E-9 | 8 | 60 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 4.02E-10 | 10 | 66 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-4 | 11 | 55 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.46E-9 | 12 | 58 | No hit | No description |
| Pfam | PF00249 | 2.1E-8 | 12 | 55 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 17.641 | 114 | 171 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.62E-16 | 118 | 172 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-17 | 119 | 169 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.2E-10 | 119 | 169 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-10 | 122 | 165 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.7E-11 | 122 | 166 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.38E-10 | 122 | 167 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 296 aa Download sequence Send to blast |
MTVDEAGSSS LWSLEQDKAF ENALASHPED DSDRWEKIAV DVPGKTVEEI KHHYELLVED 60 VNLIESGCVP LPSYSSSSDG SANHAGDEGT TKKNGHFGNC NGDSNHGSKT SRSDQEHRKG 120 IAWTEDEHRL FLLGLDKYGK GDWRSISRNF VVTRTPTQVA SHAQKYFIRL NSMNKERRRS 180 SIHDITSVAN GDISATQGPI TGQANGSGAA PSGKPTKQPP QSAAGPPGVG MYGGPTMGQP 240 VGGPLVSAVG TPVSLPAPAH MAYGVRAPVP GTVVPGAPVN MGPMTYPMPP TSAHR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 1e-17 | 7 | 77 | 6 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
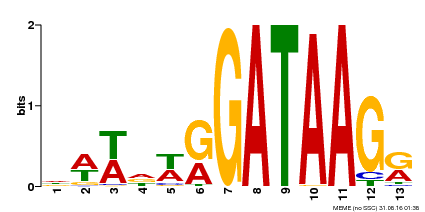
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00500 | DAP | Transfer from AT5G08520 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681895 | 0.0 | LN681895.1 Cucumis melo genomic scaffold, anchoredscaffold00005. | |||
| GenBank | LN713263 | 0.0 | LN713263.1 Cucumis melo genomic chromosome, chr_9. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008465139.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Refseq | XP_008465150.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Refseq | XP_008465157.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Refseq | XP_008465164.1 | 0.0 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Swissprot | Q9FNN6 | 1e-127 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A1S3CNM4 | 0.0 | A0A1S3CNM4_CUCME; transcription factor DIVARICATA-like | ||||
| STRING | XP_008465139.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4694 | 33 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G08520.1 | 1e-114 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



