 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C002327P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1581aa MW: 177063 Da PI: 8.6712 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.7 | 0.00078 | 1547 | 1573 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
MELO3C002327P1 1547 YVCAepGCGQTFRFVSDFSRHKRKtgH 1573
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 5.5E-17 | 23 | 64 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.284 | 24 | 65 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 1.2E-13 | 25 | 58 | IPR003349 | JmjN domain |
| SMART | SM00558 | 7.8E-48 | 196 | 365 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.261 | 199 | 365 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 3.57E-25 | 210 | 378 | No hit | No description |
| Pfam | PF02373 | 2.1E-36 | 229 | 348 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 2.6E-4 | 1456 | 1486 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 17 | 1464 | 1486 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.136 | 1487 | 1516 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.072 | 1487 | 1511 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.9E-4 | 1488 | 1515 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1489 | 1511 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.64E-13 | 1498 | 1552 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.7E-8 | 1516 | 1541 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1517 | 1546 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1517 | 1541 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1519 | 1541 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.4E-9 | 1542 | 1570 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.032 | 1547 | 1578 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1547 | 1573 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1549 | 1573 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1581 aa Download sequence Send to blast |
MAGTAMAAEP TQEVLSWLKT LPLAPEYHPT LAEFQDPISY IFKIEKEASK FGICKIVPPV 60 PPSPKKTVIV NFNKSLAARA APCSDSTNSK SPPTFTTRQQ QIGFCPRKTR PVQKSVWQSG 120 EYYTFQQFEA KAKNFEKSYL KKCTRKGGLS PLEIETLYWR ATLDKPFSVE YANDMPGSAF 180 VPVSAKMFRE AGEGTTLGET AWNMRGVSRA KGSLLKFMKE EIPGVTSPMV YVAMMFSWFA 240 WHVEDHDLHS LNYLHMGAGK TWYGVPRDAA VAFEEVVRVQ GYGGEINPLV TFAVLGEKTT 300 VMSPEVLVSA GVPCCRLVQN AGEFVVTFPR AYHTGFSHGF NCGEAANIAT PEWLNVAKDA 360 AIRRASINYP PMVSHYQLLY DLALSSRAPL CSGAEPRSSR LKDKRRSEGD TVIKELFVQN 420 IVENNSLLDN LGGGASVVLL PPGSLESIYS RLRVGSHLRS KPRFPTGVCS SKEETKSPQS 480 FDYDNLALEN SPGINRVKGF YSANGPYSTL SERSTDNLCA SSSRPLNANN ERGGNVQSNG 540 LSDQRLFSCV TCGILSFACV AIIQPREQAA RYLMSADCSF FNDWVVGSGI ASEGISTKDR 600 HPVSSQQISN SGKRDKCVSD GLYDIPVLAV NRQLQLAGKS YEADLNTEKR NETSALGMLA 660 LTYGHSSDSE DDNAEADAVL NVDDAKLMIC SSEEQYQFEN SGLTSSEYSK NTAILNHDPS 720 SFGVNSADHM QFQVNDYEEF RRADSKDSFN CSSESEMDGI GSTKKNGLST RYQDSHVNGR 780 SSLDADTEKP VFDKSTETVE TENMPFAPDI DEDSSRLHVF CLEHAKEVEQ QLRPIGGVHI 840 LLLCHPVSSD YYAKLENFAA SNIACFMKNL LDYPKMEAEA KLVAQELSMS HLWTDTIFRD 900 ATQDEEKRIQ LALDCEEAIP GNGDWAVKLG INLFYSANLS HSPLYSKQMP YNSVIYNAFG 960 RSTSANSSGK PKVYQRRTGK LKRVVAGKWC GKVWMSNQVH PLLAKRDPQE EDVDIFPSWT 1020 MSDEKVDRKS ANIQKIETVK VNRKSAGKRK MNYGRGTTKK AKLVESEDMV SDASVEDCIH 1080 RHHSILRNKQ CKFVESNDPM SDDSVEDDSS RKHGVPVSKG TPYFVTDDTG SDDSLGDRHT 1140 PHRGFSGFKL PRWGEIEPSV SDDSLEHYSS QHRGKNIKSR TEKYIERQDT LSDECLESGS 1200 LKQYRRIPKS KQTKVFKKNA ISHDIRDDSF LWHHQRPSRI KKAKFIESED AVSEHSLENN 1260 SHQHRSMPQI KPAKHTAWED AFSDGPDEDD NSLLHHRNVR SNMQFREITS DDQLDDSANQ 1320 CSRRVLRRKP VKTETISQMK QEILRPAKRG ASQTLKEEFA QSLKRGGRHS LKLETPQPKI 1380 QHATNRRGKQ TKRNGKSTDL ESEEDQLGGP STRLRKRTPK PTQLSEAKVK DKKPVAKKKM 1440 KTGSSLKTPA GHRDSKARDE ESEYLCDIEG CNMSFGTKQE LALHKRNICP VKGCVKKFFS 1500 HKYLVQHRRV HMDDRPLKCP WKGCKMTFKW AWARTEHIRV HTGARPYVCA EPGCGQTFRF 1560 VSDFSRHKRK TGHSTKKGRG * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-76 | 15 | 379 | 6 | 346 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-76 | 15 | 379 | 6 | 346 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1414 | 1440 | RKRTPKPTQLSEAKVKDKKPVAKKKMK |
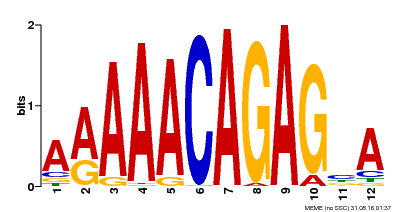
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681932 | 0.0 | LN681932.1 Cucumis melo genomic scaffold, anchoredscaffold00001. | |||
| GenBank | LN713266 | 0.0 | LN713266.1 Cucumis melo genomic chromosome, chr_12. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008439230.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A1S3AXW7 | 0.0 | A0A1S3AXW7_CUCME; lysine-specific demethylase REF6 | ||||
| STRING | XP_008439230.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



