 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000286804 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 1389aa MW: 158262 Da PI: 5.4555 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 87.5 | 2e-27 | 76 | 163 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++e+++++f Cs++g + e+++ ++r+ ++++t+Cka+++vk++ dgkw ++++ +eHnHel p
MDP0000286804 76 SFYQEYAKSMGFTTSIKNSRRSKKSKEFIDAKFACSRYGVTPESDSG---TSRRPTVKKTDCKASMHVKRRADGKWIIHEFIKEHNHELLP 163
5****************************************999888...7788889*******************************975 PP
| |||||||
| 2 | FAR1 | 26 | 3e-08 | 1057 | 1111 | 2 | 57 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtrae 57
+Y+eYA +vGF + ++ s++sk++g++++ + Cs+ g++++ + ++ r++++
MDP0000286804 1057 YYREYAWSVGFGIIIKVSRRSKKSGKFIDIKIACSRFGSKHDFIYA-TSGRNKKPA 1111
9**************************************9987766.555555554 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 6.7E-25 | 76 | 163 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.3E-22 | 283 | 375 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 10.175 | 563 | 599 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 3.6E-5 | 573 | 598 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.5E-7 | 574 | 601 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF03101 | 2.3E-6 | 1057 | 1104 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.1E-23 | 1166 | 1256 | IPR018289 | MULE transposase domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1389 aa Download sequence Send to blast |
MDRNQNSGET VVDGPEKMSV GAVNENMNMV AVAEEVQNRG GGVITSPEKD VQVFEGEADF 60 EPCNGIEFES HEAAYSFYQE YAKSMGFTTS IKNSRRSKKS KEFIDAKFAC SRYGVTPESD 120 SGTSRRPTVK KTDCKASMHV KRRADGKWII HEFIKEHNHE LLPALAYHFR IHRNVKLAEK 180 NNVDILHAVS ERTRKMYVEM SRHSGGYQST GFPRADTNYQ FDKCRDLSLD LGDAQVMLEY 240 FKRIQKENPN FFYAIDLNEE RRLRNLFWVD AKSRSDYKSF NDVVSFDTSY IKINDKLPIA 300 PFVGVNNHFQ SMLLGCALVA DDTKSTFVWL LKTWLRAMGG QGPKVIITDQ DQTLKAAIGE 360 VFPDARHCFS LWSVLEKIPE TLAHVIKRHD NFLSKFNECI FKSSTDEQFV FRWWEMVNSF 420 ELQDDEWIRL LYDDRNRWVP TYIGDTFLAG MCTPQRSESM NSFFDKYIHR KISLREFVKQ 480 YGTILQNRYE EESIADFDTW HKQPALKSPS PWEKQMSTVY THAVFKKFQV EVLGVVGCQP 540 KKEHEDGPTT TFRVQDCEKD EYFMVAWNET KSEVSCSCHL FEYKGFLCRH TLIVLQMCGL 600 SSIPSHYILK RWTKDAKIRQ SMVEETERVQ TRVQRYNDLC KRAIELSEEG SSSEETYNIA 660 LRTLVEALKN CVNVNNSNGT VIEFSGTVHG IREAEEENQG SPAAKAIRKR NTNRKRKVPA 720 KQDGVLVESQ DSLQQMDNLS SDGIALNGYY GAQQNVHGLV QLNLMEPPHD SYYVNQQSMQ 780 GLGQLNSIAP NHDGFFGAQQ SIHGMGQLDF RPSSSFSYSL QICSADRYLE CTNQCLVSED 840 IEVHGLEVCK SILCNAKQRD LELLSCEQDQ LAIRSDKDVN IVDVTDEINV DAEPSSPIAS 900 DRVKEVDGFS AEVSVMSDKD VNVVDAPYEI IVEVNVSSQT TGGHFKEVDG LNAEQSVVSD 960 RDVDIVDVND EINVDKEHVS SPTSDRVKEV DGLNADKSVE SDKDVDIVDI TEQCVSSLND 1020 QSDLNNVGVD SLDKGKGVVE EPENGLEFES KEEAYSYYRE YAWSVGFGII IKVSRRSKKS 1080 GKFIDIKIAC SRFGSKHDFI YATSGRNKKP ATVVPQKTGL QLALDEEDDE DPNFFYAIDF 1140 DHEKRLRSVF WVDSKCKHDY SSFCDVVFFD TYYVSNNYKI PFVPIVGVNH HCLYILLGCA 1200 LIGEETTSAF IWLKQTWLRA VGGQAPGVVI TDQEKYLEEA VEDVFTDARH CFCLWHDLFL 1260 VGLSTKERSG SKTSFFDGYI SQEATFKDFV EQRFHYFNVN NSVRSVSEPS VSANHGFNDV 1320 EVVNPSSSVA KLSNRKKTYK KRKELMKSRA HEIDNCHVPQ QESERLVHFL YASVERQFGK 1380 TISLRPIFT |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
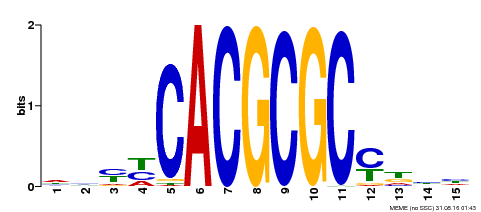
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008378343.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 isoform X1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A251Q247 | 0.0 | A0A251Q247_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008378343.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000286804 |



