 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MDP0000216756 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Malus
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 863aa MW: 99099.4 Da PI: 6.4236 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 85.2 | 1.1e-26 | 104 | 206 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk............tekerr....traetrtgCkaklkvkkekdgkwevtk 80
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ + +r+ ++t+Cka+++vk++ dgkw++++
MDP0000216756 104 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYDKSynrprarqnkqdP----EnptgRRSCSKTDCKASMHVKRRPDGKWVIHN 195
5******************************************99999999977655443....144559999*********************** PP
FAR1 81 leleHnHelap 91
+++eHnHel p
MDP0000216756 196 FVKEHNHELLP 206
********975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.5E-24 | 104 | 206 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.8E-31 | 304 | 396 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.025 | 584 | 620 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.1E-6 | 595 | 622 | IPR006564 | Zinc finger, PMZ-type |
| Pfam | PF04434 | 6.3E-5 | 595 | 618 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 863 aa Download sequence Send to blast |
MMFSDSNQLK LNNLTMDIDL RLPSGEHDKE DEEPHEIDNM LEHEEKVHNG DIENGNIEDV 60 GDEVLAEDGG DLNSPTPDIV VFKEDTNLEP LFGMEFESHG EAYSFYQEYA RSMGFNTAIQ 120 NSRRSKTSRE FIDAKFACSR YGTKREYDKS YNRPRARQNK QDPENPTGRR SCSKTDCKAS 180 MHVKRRPDGK WVIHNFVKEH NHELLPAQAV SEQTRKMYAA MARQFAEYKN VVGLKSDSKN 240 PFDKGRNLAL EAGDLKILLE FFTQMQNMNS NFFYSIDLGE DQRLKSLFWV DAKSRHDYIN 300 FSDIVSFDTT YIRNKYKMPL ALFVGVNQHY QFVLLGCALV SDESTATFSW LMQTWLKAMG 360 GQAPKVIITD HDKSIKSVIA EVFPSAYHCF SLWHILGKVS ENLGHVIKQH ENFMAKFEKC 420 IHRSSTNEEF EKRWWKIVEK FELKEDEWTQ LLYEDRKQWV PXYMREICLA GMSAVQRSES 480 VNSFFDKYVH KKTTVQEFLK QYEAILQDRY DEEAKADYDT WNKPPTLRSP SPLEKSVSGL 540 YTHAVFKKIQ GEVLGAVACH PKRERQDETS IIFNVVDFEK NQDFIVTWNE MKTEVSCVCC 600 LFEYKGYLCR HALIVLQICG LSAIPSQYVL KRWTKDAKSQ HFVGEESDIV LSRVQKFNDL 660 CQRAMKVIEE GTLSQESYSV ACRALEEAFG NCVSVNNSSK SLLEASTSVT HGLLCIEDDN 720 QNRSMSSKTN KKKNPTKKRK VNCEPEVMTV GAQDGLQQME KLTSRTVTLD GYYGAQPNVQ 780 GMVQLNLMAP TRDNYYGNQQ TIQGLGQLNS IAPSHDGYYS AQQSMHGLGQ MDFFRTPTGF 840 AYGIRDDPNV RTTPLHEDAS RHA |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mdo.24731 | 1e-105 | fruit | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
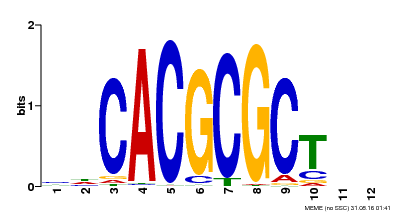
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008368542.2 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3-like isoform X1 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A251Q297 | 0.0 | A0A251Q297_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008368542.1 | 0.0 | (Malus domestica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF40 | 30 | 530 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | MDP0000216756 |



