 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10040945 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 359aa MW: 39158.3 Da PI: 7.2218 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99 | 3.3e-31 | 80 | 133 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +F++av+ LGG+++AtPk +lelm++kgL+++hvkSHLQ+YR+
Lus10040945 80 PRLRWTPDLHLCFINAVDTLGGQDRATPKLVLELMNIKGLSIAHVKSHLQMYRS 133
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.997 | 76 | 136 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.3E-29 | 76 | 134 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.91E-14 | 78 | 134 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-23 | 80 | 135 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.5E-8 | 81 | 132 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 359 aa Download sequence Send to blast |
MGSSSRHPDG NSSKSSFLSN TDAFTSEDEQ EEEEDEEGEA GSDCDVISTN SSVDDEEKKK 60 AAAGNSSGGC VRKYNRSKMP RLRWTPDLHL CFINAVDTLG GQDRATPKLV LELMNIKGLS 120 IAHVKSHLQM YRSKKLDDPD QGQHQGLLLQ GKDTHVYNLS QLPMLRTFNQ TTSSTPFRYG 180 DGNGRSGHQM YGKHYIGEGG GGGGVNRSKQ GSPYYGYKQN PFGTSSSQVS IHRLTHHHLL 240 QVSSKLLSRP SWPAQITKPI TTTMESTKTF GNHNNNDGIL KRKTREPDDD DHLNDLDLNL 300 SLKVKAAKIV DEEAAAAPTS RILQKEVDQG STTTSLSLSL SSSSKCTRMA ASTLDLTL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 8e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-18 | 81 | 135 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
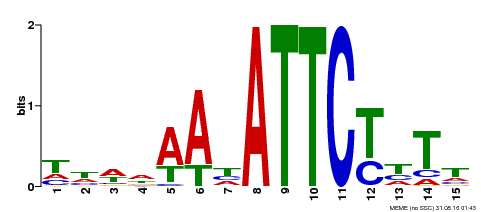
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10040945 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028084429.1 | 6e-66 | myb family transcription factor PHL8 isoform X2 | ||||
| TrEMBL | A0A067LJD0 | 2e-57 | A0A067LJD0_JATCU; Uncharacterized protein | ||||
| STRING | Lus10040945 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 4e-40 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10040945 |



