 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lus10039883 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Linaceae; Linum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 458aa MW: 49770.9 Da PI: 6.9916 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 88.5 | 6.1e-28 | 177 | 231 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G +kA+P++ilelm++++Lt+++v+SHLQkYR++
Lus10039883 177 KMKVDWTPELHRRFVQAVEQL-GVDKAVPSRILELMGIHCLTRHNVASHLQKYRSH 231
5689*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.981 | 174 | 233 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.34E-18 | 175 | 234 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.9E-27 | 175 | 234 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.6E-26 | 177 | 231 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.0E-8 | 180 | 229 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 458 aa Download sequence Send to blast |
MLAVSVAAPL GVDESSFQQG KMMEEVNNFS DGFFCDGNLL LMESIDFDDL FVGMDDGDVL 60 PDLEMDSEIL TDFSQESEVI IHDGATTTAT GFETVGNEGE DVVIIIEDGK DSNKDGMGTG 120 KEEVGQAASP SSSASMVMNP PGSDGVRSGG EKRKKSSTAS SHQSNIGGKN NNQGKRKMKV 180 DWTPELHRRF VQAVEQLGVD KAVPSRILEL MGIHCLTRHN VASHLQKYRS HRKHILAREA 240 EAASWSHRRQ MYGIAGVTSH SNSVDVNHQN HHRRPWKHAP ATVGFPPMHH HLVRPLHVWG 300 HPHQHQPLWH HNRRVGATGE GVTPSPGIAS FPQAAAPAAV RPRRLNYDHH QRVLSLSHSQ 360 KFTTPPILSI IPPPHAMYKP GHGVSPPATS ASSPNQPFDF QPSNESIDAA IGDVLSKPWL 420 PLPLGLKPPA VDTVLVELHC QGISNVPPRP STTSFGK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 7e-15 | 174 | 235 | 2 | 63 | ARR10-B |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 152 | 178 | RKKSSTASSHQSNIGGKNNNQGKRKMK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
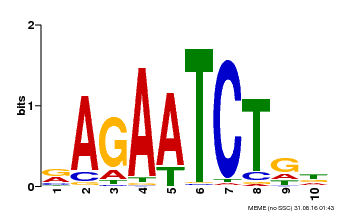
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lus10039883 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| STRING | Lus10039883 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4242 | 34 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 1e-68 | GBF's pro-rich region-interacting factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Lus10039883 |



