 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj2g3v0621850.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 463aa MW: 50452.5 Da PI: 5.5762 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52 | 1.6e-16 | 58 | 102 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++++a k++G g W+ I +++g +t+ q++s+ qk+
Lj2g3v0621850.2 58 REKWTEEEHQKFLEALKLYGRG-WRQIEEHIG-SKTAVQIRSHAQKF 102
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.68E-16 | 52 | 106 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.831 | 53 | 107 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.5E-10 | 55 | 106 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.0E-16 | 56 | 105 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 7.3E-13 | 57 | 105 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.7E-14 | 58 | 101 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.38E-11 | 60 | 103 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 463 aa Download sequence Send to blast |
MEMKDQVEGP RSTIYGTASN FPSNGGTQSD NVAKLPDTPA GNSLAPKARK PYTITKQREK 60 WTEEEHQKFL EALKLYGRGW RQIEEHIGSK TAVQIRSHAQ KFFSKVVRES DGSAESSIQP 120 IAIPPPRPKR KPLHPYPRKS ADSFKGQTVP NEPEKSPSAN LSGGEKETQS PTSVLSAFGS 180 EAFGSAFSEQ TNRCLSPNSC TTDIHSMSLS PAEKDNDCMT SKPAEVEEKG SLASVNLTTG 240 LNPLMCMKSE IGAEETEGLK EDATNMPPIS SSIKLFGRTV SMVGNLMSMK VDDENIKPET 300 IEMDDVENVK VGQVGASEPL DTQLSLGLSC GKLTSIEHPK ENLCSSEDDP DSSLPSLSLY 360 QGLSAFNLRP CNHEILNPMP LRPCLKVRAR EEESSCTGSN TESVGKNSDT VDSKLQKYHE 420 DGVAPQKSGR GFVPYKRCLA ERDANSLIVG LEEREGQRAR LCS |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.5761 | 0.0 | cell culture| flower | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
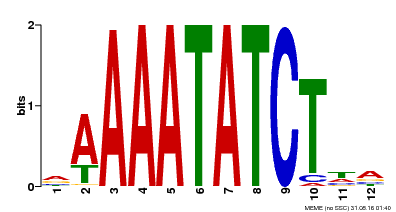
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj2g3v0621850.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015044 | 6e-39 | AP015044.1 Vigna angularis var. angularis DNA, chromosome 11, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003548314.1 | 0.0 | protein REVEILLE 7 | ||||
| Refseq | XP_014624432.1 | 0.0 | protein REVEILLE 7 | ||||
| Refseq | XP_028207255.1 | 0.0 | protein REVEILLE 7-like | ||||
| Refseq | XP_028207256.1 | 0.0 | protein REVEILLE 7-like | ||||
| TrEMBL | A0A371HX55 | 0.0 | A0A371HX55_MUCPR; Protein REVEILLE 7 (Fragment) | ||||
| TrEMBL | A0A445GLJ1 | 0.0 | A0A445GLJ1_GLYSO; Protein REVEILLE 7 isoform A | ||||
| TrEMBL | I1MQL7 | 0.0 | I1MQL7_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA16G34340.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7313 | 32 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 1e-53 | MYB_related family protein | ||||



