 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj1g3v0416400.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 292aa MW: 32082 Da PI: 8.0267 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43.3 | 8.2e-14 | 76 | 120 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE++l++ + ++ G+g+W+ I++ + k+Rt q+ s+ qky
Lj1g3v0416400.1 76 PWTEEEHKLFLLGLQKVGKGDWRGISKNFVKTRTSTQVASHAQKY 120
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.266 | 68 | 125 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.2E-16 | 71 | 125 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 8.4E-17 | 72 | 123 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 4.2E-10 | 73 | 123 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-11 | 75 | 120 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.92E-9 | 76 | 121 | No hit | No description |
| Pfam | PF00249 | 2.0E-11 | 76 | 120 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 292 aa Download sequence Send to blast |
MSNTAVSGAG EFMLFGVRVT VDSMRKCVSM NNLSQYEHPH DAPNNKEHAA GYASADEAPA 60 PLPVKLANPD RKRGVPWTEE EHKLFLLGLQ KVGKGDWRGI SKNFVKTRTS TQVASHAQKY 120 FLRRANLTRR RRRSSLFDIT TDTVSAAQME EEQIQHQDNM SHSCPIYPAT ATETPESSNR 180 NGYPMMPMYS MAVGSGVNSV LARNQMEKLT LGQENVEQNE VSKLACTIPA VPDHKASSSV 240 SDISSNTSSI VDPPTLSLGL SLSSDQRQKS PRHTPLHAVP CFNNEDSIIT VA |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Lja.11196 | 0.0 | pod| root | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds selectively to the DNA sequence 5'-[GA]GATAA-3' and may act as a transcription factor involved in the regulation of drought-responsive genes. Enhances stomatal closure in response to abscisic acid (ABA). Confers drought and salt tolerance. {ECO:0000269|PubMed:21030505}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00631 | PBM | Transfer from PK23829.1 | Download |
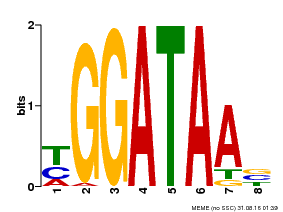
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj1g3v0416400.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, high salinity, and ABA. {ECO:0000269|PubMed:21030505}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP009643 | 0.0 | AP009643.1 Lotus japonicus genomic DNA, clone: LjB09A03, BM2001, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027329958.1 | 1e-122 | transcription factor MYB1R1-like | ||||
| Swissprot | Q2V9B0 | 6e-58 | MY1R1_SOLTU; Transcription factor MYB1R1 | ||||
| TrEMBL | A0A371F4T0 | 1e-112 | A0A371F4T0_MUCPR; Uncharacterized protein (Fragment) | ||||
| STRING | GLYMA04G34720.1 | 1e-109 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4248 | 31 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G74840.2 | 2e-57 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



