 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj0g3v0320969.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 318aa MW: 36712.2 Da PI: 7.3316 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 175.5 | 1.5e-54 | 10 | 139 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvka.eekewyfFskrdkkyatgkrknratksgyWkatgkdke 91
+ppGfrFhPtdeel+ +yL+kkv+ + ++l +vi+evd++k+ePwdL+ k ++ + ++ewyfFs++dkky+tg+r+nrat++g+Wkatg+dk+
Lj0g3v0320969.2 10 VPPGFRFHPTDEELLYYYLRKKVSYEAIDL-DVIREVDLNKLEPWDLKdKcRIGSgPQNEWYFFSHKDKKYPTGTRTNRATTAGFWKATGRDKA 102
69****************************.9***************943444443456*********************************** PP
NAM 92 vlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
+++++++ +g++ktLvfy+grap+g+ktdW+mheyrl
Lj0g3v0320969.2 103 IYHSNSKRIGMRKTLVFYTGRAPHGQKTDWIMHEYRL 139
***********************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.35E-60 | 5 | 159 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 58.612 | 10 | 159 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.9E-28 | 11 | 139 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003002 | Biological Process | regionalization | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009834 | Biological Process | plant-type secondary cell wall biogenesis | ||||
| GO:0010455 | Biological Process | positive regulation of cell fate commitment | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 318 aa Download sequence Send to blast |
MMQGSGQLAV PPGFRFHPTD EELLYYYLRK KVSYEAIDLD VIREVDLNKL EPWDLKDKCR 60 IGSGPQNEWY FFSHKDKKYP TGTRTNRATT AGFWKATGRD KAIYHSNSKR IGMRKTLVFY 120 TGRAPHGQKT DWIMHEYRLD DDDAEVQEDG WVVCRVFKKK NTTRGFQQQE IEEEEHFSHM 180 MRPTNNGPCH QILESKHHHH NMQGLYDSSS FDGTIHLPQL FSPESAAVAP TNCMNTMDIL 240 ECSQNLLRLT TTTTGCGLNL MQQQQGDQRF TGDWSFLDKL LASHHPTQPH HAVGSTSAQK 300 FPFHYLGNEA HDIMKFSK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 5e-52 | 7 | 164 | 14 | 170 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 5e-52 | 7 | 164 | 14 | 170 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 5e-52 | 7 | 164 | 14 | 170 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 5e-52 | 7 | 164 | 14 | 170 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 5e-52 | 7 | 164 | 14 | 170 | NAC domain-containing protein 19 |
| 4dul_B | 5e-52 | 7 | 164 | 14 | 170 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: First expressed at early heart stage onward in all root basal daughter cells resulting from horizontal divisions in the COL progenitors and is later maintained in these cells. Present in root stem cell daughters and accumulates in maturing root cap layers. Detectable from very early stages of lateral root development. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Uniprot | TISSUE SPECIFICITY: Accumulates in maturing root cap cells, in both COL and LRC cells. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription regulator. Together with BRN1 and BRN2, regulates cellular maturation of root cap. Represses stem cell-like divisions in the root cap daughter cells, and thus promotes daughter cell fate. Inhibits expression of its positive regulator FEZ in a feedback loop for controlled switches in cell division plane. Promotes the expression of genes involved in secondary cell walls (SCW) biosynthesis. {ECO:0000269|PubMed:19081078, ECO:0000269|PubMed:20197506}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00250 | DAP | Transfer from AT1G79580 | Download |
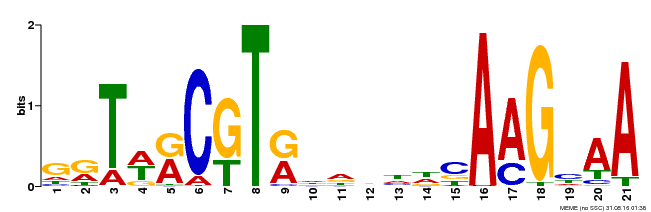
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj0g3v0320969.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By FEZ in oriented-divised root cap stem cells. {ECO:0000269|PubMed:19081078}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027343937.1 | 0.0 | protein SOMBRERO | ||||
| Swissprot | Q9MA17 | 1e-119 | SMB_ARATH; Protein SOMBRERO | ||||
| TrEMBL | A0A1J7HFW0 | 0.0 | A0A1J7HFW0_LUPAN; Uncharacterized protein | ||||
| STRING | GLYMA08G17140.1 | 1e-180 | (Glycine max) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G79580.3 | 1e-106 | NAC family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



