|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
LPERR08G12050.1 |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
| Family |
HD-ZIP |
| Protein Properties |
Length: 334aa MW: 35901.2 Da PI: 4.2642 |
| Description |
HD-ZIP family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| LPERR08G12050.1 | genome | OGE | View CDS |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Homeobox | 62.7 | 5.4e-20 | 82 | 135 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++t+eq++ Le+ Fe+++++ e+++eLA++lg+ rqV vWFqNrRa++k
LPERR08G12050.1 82 KKRRLTAEQVQMLERSFEEENKLEPERKTELARRLGMAPRQVAVWFQNRRARWK 135
56789************************************************9 PP
|
| 2 | HD-ZIP_I/II | 127.7 | 5.1e-41 | 81 | 170 | 1 | 90 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLree 90
ekkrrl++eqv++LE+sFeee+kLeperK+elar+Lg+ prqvavWFqnrRAR+ktkqlE+d+++Lk+a+dal++++++L +++++Lr++
LPERR08G12050.1 81 EKKRRLTAEQVQMLERSFEEENKLEPERKTELARRLGMAPRQVAVWFQNRRARWKTKQLETDFDRLKAAFDALAADHQALLDDNHRLRAQ 170
69*************************************************************************************975 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0001558 | Biological Process | regulation of cell growth |
| GO:0009637 | Biological Process | response to blue light |
| GO:0009651 | Biological Process | response to salt stress |
| GO:0009965 | Biological Process | leaf morphogenesis |
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated |
| GO:0005634 | Cellular Component | nucleus |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0042803 | Molecular Function | protein homodimerization activity |
| GO:0043565 | Molecular Function | sequence-specific DNA binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 334 aa
Download sequence Send
to blast |
MDPGRVVFDS GVARRAGGAG GGGGGAQMLL FGGGGSANSG GFFRGVPTAV MGMEAGTKRP 60
FFTTHEEVIV EEEYYDEQAP EKKRRLTAEQ VQMLERSFEE ENKLEPERKT ELARRLGMAP 120
RQVAVWFQNR RARWKTKQLE TDFDRLKAAF DALAADHQAL LDDNHRLRAQ ETSRPSSAII 180
TTAAQEVDQP DEHTEAASDT GCATVDDALA PPLATARQQQ QLKDEFLSSG ATNNDDRSSG 240
GGAAVVVFDA TEGTNDVSCE SAYFAAAAEA YERDCAGQYP LSSEEEDTGA VSDEGCSFDL 300
PDDAMFGAAA GIVHHAAAGD EEAQLGSWTA WFWS
|
| Nucleic Localization
Signal ? help
Back to Top |
 |
| No. |
Start |
End |
Sequence |
| 1 | 129 | 137 | RRARWKTKQ |
| Functional Description ? help
Back to Top |
| Source |
Description |
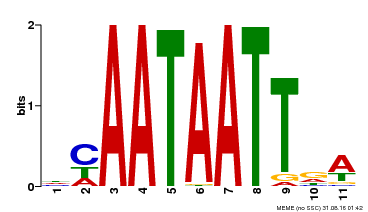
| UniProt | Probable transcription activator that binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. {ECO:0000269|PubMed:10732669}. |
| UniProt | Probable transcription activator that binds to the DNA sequence 5'-CAAT[AT]ATTG-3'. {ECO:0000269|PubMed:10732669}. |
| Annotation --
Nucleotide ? help
Back to Top |
| Source |
Hit ID |
E-value |
Description |
| GenBank | AF145729 | 1e-135 | AF145729.1 Oryza sativa homeodomain leucine zipper protein (Oshox5) mRNA, complete cds. |
| GenBank | AK100634 | 1e-135 | AK100634.1 Oryza sativa Japonica Group cDNA clone:J023110A12, full insert sequence. |
| GenBank | AK101569 | 1e-135 | AK101569.1 Oryza sativa Japonica Group cDNA clone:J033050G15, full insert sequence. |
| GenBank | AP004667 | 1e-135 | AP004667.3 Oryza sativa Japonica Group genomic DNA, chromosome 8, PAC clone:P0433E10. |
| GenBank | AP014964 | 1e-135 | AP014964.1 Oryza sativa Japonica Group DNA, chromosome 8, cultivar: Nipponbare, complete sequence. |
| GenBank | CP012616 | 1e-135 | CP012616.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 8 sequence. |