 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR05G01400.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 374aa MW: 42566.6 Da PI: 9.021 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 19.6 | 2.4e-06 | 77 | 97 | 3 | 23 |
ETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 CpdCgksFsrksnLkrHirtH 23
C+ Cg +F+ + +L++H+++H
LPERR05G01400.1 77 CKVCGARFKKPAHLRQHMQSH 97
*******************99 PP
| |||||||
| 2 | zf-C2H2 | 19.7 | 2.2e-06 | 103 | 127 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ C+ s+srk++L+rH+ tH
LPERR05G01400.1 103 FSCHvdGCPFSYSRKDHLNRHLLTH 127
89*********************99 PP
| |||||||
| 3 | zf-C2H2 | 18.2 | 6.8e-06 | 132 | 157 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+ Cp C++ Fs+k n++rH + H
LPERR05G01400.1 132 FACPmeGCNRKFSTKGNMQRHVQEmH 157
78*******************99877 PP
| |||||||
| 4 | zf-C2H2 | 20.6 | 1.1e-06 | 169 | 193 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y Cp Cgk+F+ s L++H +H
LPERR05G01400.1 169 YICPeaSCGKTFKYASKLQKHEESH 193
89*******************9988 PP
| |||||||
| 5 | zf-C2H2 | 14.9 | 7.7e-05 | 260 | 284 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ dC+ sFs ksnL +H + H
LPERR05G01400.1 260 KCHfeDCKCSFSKKSNLDKHVKAvH 284
7*******************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 27 | 31 | 53 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.986 | 75 | 102 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.013 | 75 | 97 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 9.2E-8 | 77 | 101 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Pfam | PF13894 | 0.0012 | 77 | 97 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 77 | 97 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 7.6E-11 | 87 | 129 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 8.5E-10 | 102 | 129 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.349 | 103 | 132 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0016 | 103 | 127 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 105 | 127 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.17E-7 | 113 | 163 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.9E-8 | 130 | 158 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.24 | 132 | 162 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.9E-5 | 132 | 157 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 134 | 157 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-6 | 165 | 193 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.15E-5 | 166 | 193 | No hit | No description |
| PROSITE profile | PS50157 | 11.427 | 169 | 193 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.003 | 169 | 193 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 171 | 193 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 18 | 201 | 226 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.517 | 202 | 226 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 203 | 226 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 25 | 229 | 250 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 2.42E-7 | 240 | 301 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.9E-5 | 240 | 281 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.988 | 259 | 289 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.13 | 259 | 284 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 261 | 284 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-5 | 283 | 311 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 8.767 | 290 | 316 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3 | 290 | 316 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 292 | 316 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 374 aa Download sequence Send to blast |
MRSVEAGAEE GESKGAAAAA AANPGRGIRR YKCEFCSVVR SKKCLIRAHV LEHHKDEVDD 60 YLRRGGGQSR KDIDRDCKVC GARFKKPAHL RQHMQSHSLE RPFSCHVDGC PFSYSRKDHL 120 NRHLLTHQGK LFACPMEGCN RKFSTKGNMQ RHVQEMHKDG SPCESKKEYI CPEASCGKTF 180 KYASKLQKHE ESHVKLDYSE VFCCEPGCMK AFTNLECLKA HNESCHRHVL CDVCGTKQLK 240 KNFKRHQRMH EGSCATERVK CHFEDCKCSF SKKSNLDKHV KAVHEQKRPF VCGFSGCGKS 300 FSYKHVKDNH EKSSAHVYIQ ANFEEIDGER PRHAGGRKRK PIPVESLMRK RVVGPDDAPA 360 CDDGTEYLRW LLSG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 8e-18 | 80 | 221 | 20 | 155 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 8e-18 | 80 | 221 | 20 | 155 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
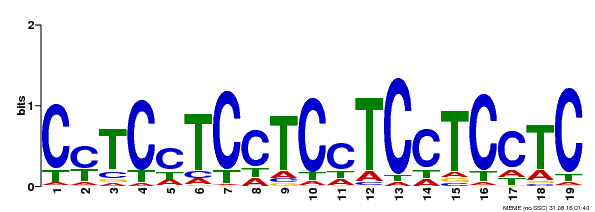
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR05G01400.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK099528 | 0.0 | AK099528.1 Oryza sativa Japonica Group cDNA clone:J013031H23, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015637660.1 | 0.0 | transcription factor IIIA isoform X1 | ||||
| Swissprot | Q84MZ4 | 1e-113 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A0D9WC64 | 0.0 | A0A0D9WC64_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR05G01400.1 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-114 | transcription factor IIIA | ||||



