 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV25396.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 457aa MW: 51622.5 Da PI: 6.122 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 46.7 | 5.6e-15 | 280 | 326 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn Er+RRdriN+++ L+el+P++ K +Ka++L +++eY+ksLq
KZV25396.1 280 VHNLSERKRRDRINEKMRALQELIPHS-----HKSDKASMLDEVIEYMKSLQ 326
5*************************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 7.72E-19 | 274 | 340 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 16.856 | 276 | 325 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 2.92E-15 | 279 | 330 | No hit | No description |
| Pfam | PF00010 | 1.9E-12 | 280 | 326 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.0E-18 | 280 | 337 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 3.8E-16 | 282 | 331 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 457 aa Download sequence Send to blast |
MERLDNELVE LLWQNGEVVL NRQTHRNSGN SSTESKQVHN HDEQKSRFAM TWTNLIQDEE 60 AISWIHCPID ESFEEELCSE FLTEIPSSNP VFEGESFLRF SGSHQHELAP ISLPPQRFET 120 LDSTRQDQIL RGDRKDGNLD LPFERREFTY MGQDDAHELS VMTIGSSHCG SNQVASSCGV 180 GSKPVSAKVD ANYVQKVSHL SDQSTETETV GQALTSSSGG SGSTFWKTAN QIDENNCHKR 240 KSRDGEQSDC LSDAIELEYG SENKSSRKSG NTRRSRVAEV HNLSERKRRD RINEKMRALQ 300 ELIPHSHKSD KASMLDEVIE YMKSLQSQIQ FMWMGSQMAQ IVVPGIPNYM SRTGMGISPI 360 AMIPQIHNLM RFSRLPQIYQ AMTVAPIPNQ TAADLRPVLK PVSYQNQMQS PSFHEQYANY 420 RNLYLMQNTS QSMNMFGLSS QSTQPNNVLA PTKYGSG |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 284 | 289 | ERKRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
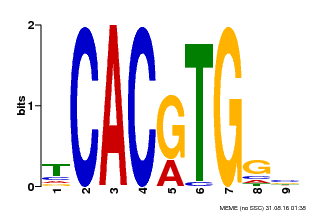
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV25396.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011100371.1 | 1e-162 | transcription factor PIF4 | ||||
| Refseq | XP_011100372.1 | 1e-162 | transcription factor PIF4 | ||||
| TrEMBL | A0A2Z7AVH4 | 0.0 | A0A2Z7AVH4_9LAMI; Uncharacterized protein | ||||
| STRING | XP_009602219.1 | 1e-124 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 3e-40 | phytochrome interacting factor 4 | ||||



