 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN41269.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 561aa MW: 64083.3 Da PI: 8.2665 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 90.6 | 1.7e-28 | 26 | 109 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW++qe+laL+++r++m++ +r+++lk+plWeevs+k++e g++rs+k+Ckek+en+ k+ k++ke++++++ + +++fdql+
KHN41269.1 26 RWPRQETLALLKIRSDMDTVFRDSSLKGPLWEEVSRKLAELGYQRSAKKCKEKFENVYKYNKRTKENKSGKSH--GKAYKFFDQLQ 109
8*****************************************************************9999754..456******98 PP
| |||||||
| 2 | trihelix | 103 | 2.3e-32 | 381 | 466 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev+aLi++r+++e++++++ k+plWe++s++m++ g++rs+k+Ckekwen+nk++k+++e++k+r e+s+tcpyf++lea
KHN41269.1 381 RWPKAEVHALIRIRTSLETKYQENGPKAPLWEDISIAMQRLGYNRSAKRCKEKWENINKYFKRVRESSKER-REDSKTCPYFHELEA 466
8*********************************************************************8.78999********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 6.922 | 19 | 83 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.13 | 23 | 85 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 2.55E-23 | 25 | 90 | No hit | No description |
| Pfam | PF13837 | 6.7E-19 | 25 | 111 | No hit | No description |
| PROSITE profile | PS50090 | 6.829 | 374 | 438 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.52 | 378 | 440 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 7.7E-22 | 380 | 467 | No hit | No description |
| CDD | cd12203 | 4.47E-25 | 380 | 445 | No hit | No description |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 561 aa Download sequence Send to blast |
MIGMDLDANS GEGNEGNNKM SFGGNRWPRQ ETLALLKIRS DMDTVFRDSS LKGPLWEEVS 60 RKLAELGYQR SAKKCKEKFE NVYKYNKRTK ENKSGKSHGK AYKFFDQLQA LENQFTVSYP 120 PKPQPTSTLT TTNTLTSDHG NKVISYVTTF PSTNPTLISP SPQTNTTTTT TTTTSTTNPR 180 DSSRPQTNNN NNSVTHSLPN MNTSFSTTTA STSSSTASDE DLEERYRRKR KWKDYFRRLT 240 RKVLLKQEEM QKKFLEAMDQ RERERVAQQD NWRMQEMARI NREHEILVQE RSTAAAKDAT 300 VIALLQKMYG QQNPTQQVEV EPPPQQKQTI PQSLPPILMP NNNFEVKKIN NGHSVTSTTT 360 GTVATAATTT TTSPVNSSSS RWPKAEVHAL IRIRTSLETK YQENGPKAPL WEDISIAMQR 420 LGYNRSAKRC KEKWENINKY FKRVRESSKE RREDSKTCPY FHELEALYKE KSKSSKNPFG 480 IFQNMKPNEM MLMMTEPLMV QPEQQWRPPP QSLEEGVGME NASEEYHENE ENGGDDDDNI 540 EEDGDSVEDE GANPCEIATN N |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
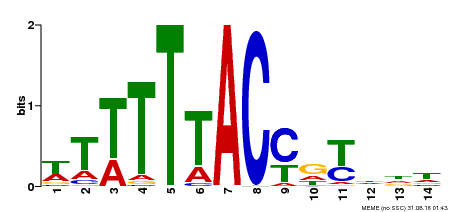
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN41269.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015044 | 1e-165 | AP015044.1 Vigna angularis var. angularis DNA, chromosome 11, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028207340.1 | 0.0 | trihelix transcription factor GT-2-like | ||||
| Swissprot | Q39117 | 1e-135 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | A0A0B2SAG9 | 0.0 | A0A0B2SAG9_GLYSO; Trihelix transcription factor GT-2 | ||||
| STRING | GLYMA16G28240.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF373 | 34 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76890.2 | 1e-117 | Trihelix family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



