 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN10328.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 536aa MW: 59329.3 Da PI: 5.5916 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 56 | 9.2e-18 | 37 | 84 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed++lvd+vk++G g+W+++ ++ g+ R++k+c++rw ++l
KHN10328.1 37 KGPWTSTEDDILVDYVKKHGEGNWNAVQKHTGLFRCGKSCRLRWANHL 84
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 46.3 | 1e-14 | 90 | 133 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++ +eE+ l++++++++G++ W++ a++++ gRt++++k++w++
KHN10328.1 90 KGPFAAEEERLIAELHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 133
78999*****************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 16.025 | 32 | 84 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.05E-29 | 36 | 131 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.8E-15 | 36 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.9E-16 | 37 | 84 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-23 | 38 | 92 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.20E-11 | 39 | 84 | No hit | No description |
| PROSITE profile | PS51294 | 24.078 | 85 | 139 | IPR017930 | Myb domain |
| SMART | SM00717 | 5.6E-15 | 89 | 137 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.1E-14 | 90 | 133 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.60E-10 | 92 | 133 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-24 | 93 | 138 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 536 aa Download sequence Send to blast |
MRRMKKDIED EVLPNDMSGA QLNDESYEGS AGIVLKKGPW TSTEDDILVD YVKKHGEGNW 60 NAVQKHTGLF RCGKSCRLRW ANHLRPNLKK GPFAAEEERL IAELHAKMGN KWARMAAHLP 120 GRTDNEIKNY WNTRIKRRQR AGLPLYPPEV SLHALQESQH SQSTGGLNGG DKMHPDFLQK 180 NSYEIHDAIF DSLKDNQGIL PYVHELSDIS VYGNKLKGLD SSQYCSFVPP TSPKRKRLKE 240 STIPFIDSCA MNKKDLYPFD QIQDNNSDKI AQSFGMQAPL DPGLSSHSSM CYSHSLSNGN 300 SSTSKPYEAM KLELPSLQYP ELDLGSWGSS PPPSLLESVD DFLQSPTAIS TLESDCSSPQ 360 NSGLLDALLY QAKTMSSSKN HCSDKSSNSS TATPGDRADS SILNVYETEW EDYADPVSPF 420 GATSILNECP ALRANANSLN GQLPVQTFTG NLPKLESVDQ VWTPNNENLT LSLLNITRPD 480 FLLASDWYDL GSGHCKNQTI TTDAATAFLG DDLATDLKHM TAGISKTMFE NPFQPL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 7e-31 | 35 | 138 | 5 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
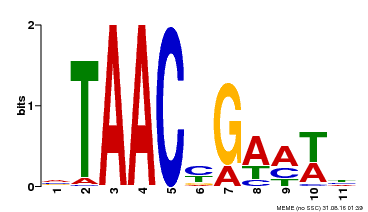
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN10328.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC525897 | 0.0 | KC525897.1 Glycine max GAMBY1-like family protein (GAMYB1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028187659.1 | 0.0 | transcription factor MYB33-like | ||||
| Refseq | XP_028187660.1 | 0.0 | transcription factor MYB33-like | ||||
| Refseq | XP_028187661.1 | 0.0 | transcription factor MYB33-like | ||||
| Refseq | XP_028187662.1 | 0.0 | transcription factor MYB33-like | ||||
| TrEMBL | A0A0B2PRI6 | 0.0 | A0A0B2PRI6_GLYSO; Transcription factor GAMYB | ||||
| STRING | GLYMA13G25716.2 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2047 | 34 | 85 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-109 | myb domain protein 33 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



