 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KFK41621.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Arabideae; Arabis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 389aa MW: 44286.2 Da PI: 7.8278 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.1 | 0.0012 | 20 | 42 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C++Cg s s k++ ++Hi +H
KFK41621.1 20 YLCQYCGISRSKKYLITSHILSH 42
56*******************96 PP
| |||||||
| 2 | zf-C2H2 | 18.6 | 5e-06 | 66 | 88 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ Cg F+ + +Lk+H+ +H
KFK41621.1 66 HNCQECGAEFKKPAHLKQHMLSH 88
67*******************99 PP
| |||||||
| 3 | zf-C2H2 | 15.5 | 5.1e-05 | 110 | 134 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C dC s++rk++L+rH+ tH
KFK41621.1 110 FTCYvdDCAASYRRKDHLNRHLLTH 134
789999*****************99 PP
| |||||||
| 4 | zf-C2H2 | 21.9 | 4.5e-07 | 191 | 215 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C+ Cgk F+ +s+L++H+ +H
KFK41621.1 191 HVCQevGCGKAFRYPSQLQKHQDSH 215
789999****************998 PP
| |||||||
| 5 | zf-C2H2 | 22.1 | 3.9e-07 | 281 | 306 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kC+ C+ +Fs +snL++H++ H
KFK41621.1 281 FKCEveGCSSTFSKPSNLQKHMKAvH 306
89********************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 25 | 20 | 42 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 6.0E-8 | 66 | 134 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 66 | 109 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0069 | 66 | 88 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.157 | 66 | 93 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 68 | 88 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.91 | 110 | 139 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.0E-9 | 110 | 136 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.11 | 110 | 134 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 112 | 134 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.24 | 139 | 164 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.808 | 139 | 169 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 141 | 164 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.03E-7 | 187 | 217 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.5E-9 | 187 | 215 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.949 | 191 | 220 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0022 | 191 | 215 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 193 | 215 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 4.8 | 223 | 248 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-4 | 225 | 246 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 225 | 248 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 25 | 251 | 272 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 8.36E-10 | 262 | 319 | No hit | No description |
| PROSITE profile | PS50157 | 13.027 | 281 | 311 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-8 | 281 | 306 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 8.9E-5 | 281 | 306 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 283 | 306 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-4 | 307 | 333 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 5.3 | 312 | 338 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 314 | 338 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 389 aa Download sequence Send to blast |
MGEESNIDMD ASLIKDIRKY LCQYCGISRS KKYLITSHIL SHHKMEMEEE RDDEACEVEE 60 EALGKHNCQE CGAEFKKPAH LKQHMLSHSI EGDLLRSSTM LASITQMRPF TCYVDDCAAS 120 YRRKDHLNRH LLTHKGKLFK CPMENCKSEF SVQGNVGRHI KEMHKNGDGS KDDIGNGDSH 180 PTEGSIGQKP HVCQEVGCGK AFRYPSQLQK HQDSHVKLDS VEAFCSEPGC MKYFTNDECL 240 KAHIRSCHQH VNCEICGSKH LKKNIKRHLR THDEDSAPGV FKCEVEGCSS TFSKPSNLQK 300 HMKAVHEDIR PFICGFPGCG MRFAYKHVRN NHENSGSHVY TCGDFVATDE NFTSRPRGGL 360 KRKQVTAEML IRKRVMPPQF ESQEEHKTC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5v3m_C | 7e-14 | 5 | 341 | 19 | 309 | Zinc finger protein 568 |
| 5wjq_D | 5e-14 | 20 | 341 | 9 | 284 | Zinc finger protein 568 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
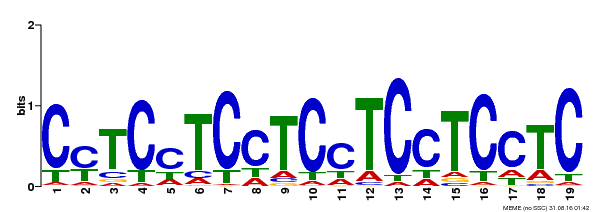
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KFK41621.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010415924.1 | 0.0 | PREDICTED: transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 0.0 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A087HHL9 | 0.0 | A0A087HHL9_ARAAL; Uncharacterized protein | ||||
| STRING | A0A087HHL9 | 0.0 | (Arabis alpina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4714 | 28 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 0.0 | transcription factor IIIA | ||||



