 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Jcr4S00401.40 | ||||||||
| Common Name | JCGZ_14339, LOC105642933 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Jatropheae; Jatropha
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 368aa MW: 40430.5 Da PI: 7.1464 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 31.7 | 3.6e-10 | 82 | 143 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT................-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg................tWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a ++ + +Wk+I + +g +t q++s+ qky
Jcr4S00401.40 82 RESWTEQEHDKFLEALQLENWEmemmgseeifafvfdrDWKKIEAFVG-SKTVIQIRSHAQKY 143
789*************988777889999********************.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.46E-12 | 76 | 149 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-5 | 80 | 149 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.4E-4 | 81 | 146 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-7 | 82 | 143 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.73E-5 | 84 | 144 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 368 aa Download sequence Send to blast |
MVSVNPNPAQ GFYFFDPMSM GLPGLNSLPP QMNSTASTTA TNNTTAAGNT ATSTTSAAAN 60 TVAFADDPNK KIRKPYTITK SRESWTEQEH DKFLEALQLE NWEMEMMGSE EIFAFVFDRD 120 WKKIEAFVGS KTVIQIRSHA QKYFLKVQKN GTSEHVPPPR PKRKAAHPYP QKAPKNVAPV 180 VTQVTGPFQP SPASLEPRYV YGSDPPSVLG NPMTGGPLTS WNFNSMAPVN ASGATKDEVG 240 LAAPANNCCY SSSNESTPRT GRTSEIIDRG DQGKAVRVMP DFAQVYSFIG SVFDPNASGH 300 LQKMKQMDPI NLETVLLLMR NLSINLTSPE FEDHRRLLAS YDIDTEKAKS VGPFVDMDTD 360 RTPKEVRG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
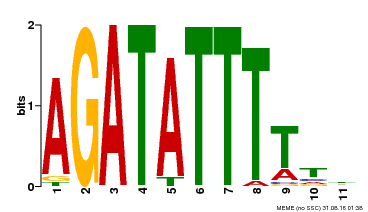
| Motif ID | Method | Source | Motif file |
| MP00423 | DAP | Transfer from AT4G01280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT093234 | 2e-41 | BT093234.1 Soybean clone JCVI-FLGm-17C23 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012083312.1 | 0.0 | protein REVEILLE 3 isoform X1 | ||||
| Swissprot | C0SVG5 | 1e-106 | RVE5_ARATH; Protein REVEILLE 5 | ||||
| TrEMBL | A0A067K0G7 | 0.0 | A0A067K0G7_JATCU; Uncharacterized protein | ||||
| STRING | cassava4.1_010350m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1119 | 33 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01280.1 | 6e-92 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 105642933 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



