 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000676.1_g00005.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1052aa MW: 119351 Da PI: 6.5179 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181.7 | 8.3e-57 | 21 | 137 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYah 87
l+ ++rwl++ ei++iL+n++k++++ e++++p++gsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah
Itr_sc000676.1_g00005.1 21 LEAQHRWLRPAEICEILKNYQKFQIAPEPPNTPPNGSLFLFDRKVLRYFRKDGHKWRKKKDGKTVKEAHERLKAGSIDVLHCYYAH 106
5679********************************************************************************** PP
CG-1 88 seenptfqrrcywlLeeelekivlvhylevk 118
+een++fqrr+yw+Leee+++ivlvhy+evk
Itr_sc000676.1_g00005.1 107 GEENENFQRRSYWMLEEEMSHIVLVHYREVK 137
****************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.963 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.6E-78 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 8.2E-50 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 5.2E-6 | 476 | 555 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.33E-16 | 477 | 563 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 2.4E-6 | 477 | 551 | IPR013783 | Immunoglobulin-like fold |
| PROSITE profile | PS50297 | 20.288 | 631 | 770 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.01E-14 | 646 | 758 | No hit | No description |
| SuperFamily | SSF48403 | 3.42E-18 | 650 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 6.0E-19 | 651 | 764 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.2E-8 | 671 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.591 | 699 | 731 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0023 | 699 | 728 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 38 | 738 | 767 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.98E-7 | 870 | 925 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 31 | 874 | 896 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 6.852 | 875 | 904 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.01 | 897 | 919 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 898 | 922 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 902 | 919 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1052 aa Download sequence Send to blast |
MATESRRYGL NAQLDIDQIL LEAQHRWLRP AEICEILKNY QKFQIAPEPP NTPPNGSLFL 60 FDRKVLRYFR KDGHKWRKKK DGKTVKEAHE RLKAGSIDVL HCYYAHGEEN ENFQRRSYWM 120 LEEEMSHIVL VHYREVKGNR VNFNHVKESQ IAIPGYQESE DTTSLNSAVA SEYEDAESVY 180 IQQKNPVAHS FLEVQSSMMQ KAQDELSVSL YPGPLANDHQ GQFSAISGIN FPSIVQGNQN 240 RNTANIYVPC QGHGFTSWKN IEETSTARYQ PALSSMNSDT ISAVQGQEDT TIGFTNNLGN 300 KQKLMNCVYA LEEWKTSEGD SLQSSEWSVD QKLNPDQACD FSTKQKPIEF CDFPTKHKHP 360 MQNDLQVQLT DADGGNFAKT KLDSDLNMRI NTDHSALKQP LLNGFLRQEL KKLDSFDRWV 420 NKELEDVNEP HMQTSSGNYW DNVGNEDGVD ESTIASQVQQ DTYVLSPSLS QDQFFSITEF 480 SPNWAYAGTE IKIRITGRFL KSQQEVDKCN WACMFGELEV PAEVISDGVL RCLTPIQKVG 540 RVPFYVTCSN RLACSEVREF EFRVNEIENA NSGSSKESLL NMRFGNLLSL ESVSFENSVP 600 GNMDYVSHLT STINSLLKED NSEWEHMFPH TSEDQLHQKF LKEKLRLWLL QKVVEGGKGP 660 NVLDESGQGV LHFAAALDYD WAIPPTLVAG VNVNFRDVNG WTALHWAAFC GRERMVVFLI 720 ALGAAPGALT DPTPQHPSGR TPADLASSNG HKGIAGYLAE ASLSTHLSSL ELKENKEDKN 780 RDASGVKAIH GVSERAATPV NDGDLPYGLS LKDSLAAVRN ATQAAARIHQ VFRVQSFQRK 840 QIKEYGDGEV GLSDERALSL LAVRKNRAGQ NNEPLHVAAM RIQNKFRSWK GRKDFLLIRQ 900 QIIKIQAHVR GHQVRKNYRK ITWSVGILEK VILRWRRKGS GLRGFKSEAL AAEGHNMQDQ 960 PKQEDDYDFL KEGRKQTEER LQKALARVQS MVQYPEARDQ YRRLLNVVSE MQETKNKYDK 1020 VLKRSGRTYD FDEDLMELEA FLNDDTFMPT AS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
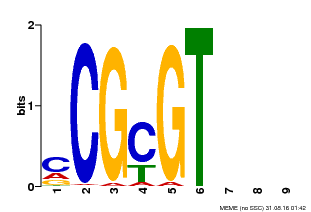
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019177324.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X2 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A1S3YKL0 | 0.0 | A0A1S3YKL0_TOBAC; Calmodulin-binding transcription activator 4 | ||||
| STRING | XP_009613615.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



