 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000654.1_g00006.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 349aa MW: 38166.2 Da PI: 6.0165 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.2 | 3.2e-32 | 95 | 148 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +Fv+ave+LGG+++AtPk +l+lm++kgL+++hvkSHLQ+YR+
Itr_sc000654.1_g00006.1 95 PRLRWTPDLHLCFVHAVERLGGQDRATPKLVLQLMNIKGLSIAHVKSHLQMYRS 148
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.55 | 91 | 151 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-30 | 91 | 149 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.33E-15 | 92 | 150 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.5E-23 | 95 | 150 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.6E-9 | 96 | 147 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MKAAETDQKV MEEMMMKEGI SCSTEDSKSG PEFLGAEIEP EGEEEEEEEE EDGKGEKGNN 60 GSSSSNSTVE ENNGKKLLGN SSGGSVRQYV RSKTPRLRWT PDLHLCFVHA VERLGGQDRA 120 TPKLVLQLMN IKGLSIAHVK SHLQMYRSKK ADEPNPVTPN GRLLLGNGDS HIFNFSQLQR 180 LQGFNPTPVT TSLRFDDSSW SHRHHHQPYS SVYNPYLNGG AAAMIRHGFG GNYVNGGRSE 240 FPMMSRQCSD NVAAPPQLFP NQRIPAAAQN GEVTAAGAGK RKLPADSDVN LDLNLSLKTR 300 DGGGGNDEKR AKGDDKYEDS TNLSLSLFPS SQNDDQTTTV KKTTLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-17 | 96 | 150 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-17 | 96 | 150 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-17 | 96 | 150 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-17 | 96 | 150 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
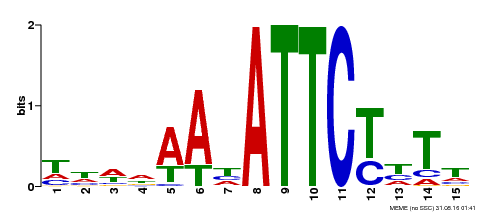
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019192424.1 | 1e-177 | PREDICTED: two-component response regulator ARR18-like | ||||
| TrEMBL | A0A328E442 | 4e-89 | A0A328E442_9ASTE; Uncharacterized protein | ||||
| STRING | XP_009791663.1 | 9e-78 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8573 | 21 | 29 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 2e-40 | G2-like family protein | ||||



