 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000357.1_g00017.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 368aa MW: 38332.2 Da PI: 9.4894 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 123.3 | 8.2e-39 | 66 | 124 | 3 | 61 |
zf-Dof 3 ekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkks 61
e+alkcprC+stntkfCy+nnysl+qPr+fCk+CrryWt+GGalrnvPvGgg+r+nk+s
Itr_sc000357.1_g00017.1 66 ETALKCPRCESTNTKFCYFNNYSLTQPRHFCKTCRRYWTRGGALRNVPVGGGCRRNKRS 124
7789*****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 2.0E-31 | 49 | 114 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 2.0E-32 | 68 | 123 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 29.544 | 69 | 123 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 71 | 107 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 368 aa Download sequence Send to blast |
MVFSSIPAYL DPANWQQLNH QVGTTTQVPS VPPAAAAPPP PHGSAGTIRP GSMADRARLA 60 NVPMPETALK CPRCESTNTK FCYFNNYSLT QPRHFCKTCR RYWTRGGALR NVPVGGGCRR 120 NKRSGKGGGG SSGGSGNNNK SPSNSSERQA SSSCSGVSTS SIMSGNSGGG LLGGPQIPPL 180 RFMSPLGQLS EHYPTGNDSI GLNINYSGLS AMAGRDNFPF QFGNNNNNNN NNNNNNNNNL 240 LGGGGGGGGL ASLLSGGIEQ WRLQTPILGG LDLSQPGLYQ FPGGSEAAPS GFLGGETSET 300 RPKFTSSSML TTQMASVKME DHNNNQESSM ARQLLGFQPG NDQWGSGGGA GWSDVSASFS 360 SSSTSNRL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence (By similarity). Probably involved in early processes for vascular development (PubMed:17583520). The PEAR proteins (e.g. DOF2.4, DOF5.1, DOF3.2, DOF1.1, DOF5.6 and DOF5.3) activate gene expression that promotes radial growth of protophloem sieve elements. Triggers the transcription of HD-ZIP III genes, especially in the central domain of vascular tissue (PubMed:30626969). {ECO:0000250|UniProtKB:Q9M2U1, ECO:0000269|PubMed:17583520, ECO:0000269|PubMed:30626969}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00295 | DAP | Transfer from AT2G37590 | Download |
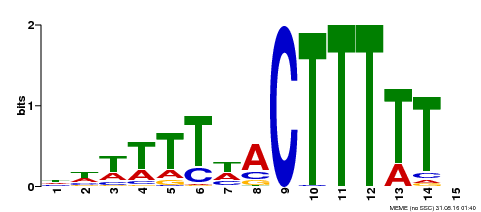
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cytokinin in procambium. Antagonized by the HD-ZIP III proteins and by mobile miR165 and miR166 microRNAs. {ECO:0000269|PubMed:30626969}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU171613 | 1e-106 | GU171613.1 Ipomoea batatas microsatellite Stv_Ipb_308 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019188085.1 | 1e-162 | PREDICTED: dof zinc finger protein DOF5.1 isoform X5 | ||||
| Swissprot | O80928 | 1e-50 | DOF24_ARATH; Dof zinc finger protein DOF2.4 | ||||
| TrEMBL | A0A068UG68 | 1e-106 | A0A068UG68_COFCA; Uncharacterized protein | ||||
| STRING | EOY07547 | 3e-95 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1917 | 24 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G37590.1 | 1e-43 | DNA binding with one finger 2.4 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



