 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000069.1_g00074.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 844aa MW: 93048.6 Da PI: 6.5043 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.1 | 7.3e-19 | 16 | 73 | 4 | 57 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
++t+eq+e+Le++++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Itr_sc000069.1_g00074.1 16 YVRYTPEQVEALERVYHECPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 73
6789****************************************************97 PP
| |||||||
| 2 | START | 172.7 | 2.4e-54 | 159 | 367 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECT CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetleviss 87
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v ++ v+e+l+d++ W ++++++++++v+ +
Itr_sc000069.1_g00074.1 159 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCTGVAARACGLVGLEPS-RVAEILKDRPSWYRDCRAVDVVNVLPT 243
789*******************************************************.8888888888****************9 PP
T..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEE CS
START 88 g..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwv 167
+ g+++l +++l+a+++l+p Rdf+ +R++ +++g++v++++S+ + q+ p+ +++vRae+lpSg+li+p+++g+s +++v
Itr_sc000069.1_g00074.1 244 AngGTIELLYMQLYAPTTLAPaRDFWLLRFTSVMEDGSLVVCERSLGNVQNGPSmppVQNFVRAEMLPSGYLIRPCDGGGSIIHIV 329
999************************************************99988899*************************** PP
E-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 168 ehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
+h+++++++++++lr+l++s+++ ++kt++a+l+++++
Itr_sc000069.1_g00074.1 330 DHMNVEAWSVPEVLRPLYESSAVLAQKTTMAALRQLRQ 367
**********************************9876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.645 | 10 | 74 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 7.8E-16 | 12 | 78 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 2.14E-17 | 14 | 78 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.48E-17 | 15 | 75 | No hit | No description |
| Pfam | PF00046 | 1.6E-16 | 16 | 73 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 8.0E-19 | 17 | 73 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 1.55E-6 | 67 | 106 | No hit | No description |
| PROSITE profile | PS50848 | 24.309 | 149 | 364 | IPR002913 | START domain |
| CDD | cd08875 | 5.77E-76 | 153 | 369 | No hit | No description |
| SMART | SM00234 | 1.4E-40 | 158 | 368 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 1.6E-22 | 158 | 365 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 4.4E-37 | 158 | 370 | No hit | No description |
| Pfam | PF01852 | 8.6E-52 | 159 | 367 | IPR002913 | START domain |
| Pfam | PF08670 | 1.5E-49 | 698 | 843 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 844 aa Download sequence Send to blast |
MSISCKDGKS AVDGKYVRYT PEQVEALERV YHECPKPSSM RRQQLIRECP ILSNIEPKQI 60 KVWFQNRRCR EKQRKEASRL QAVNRKLTAM NKLLMEENDR LQKQVSQLVY ENSYFRRQTQ 120 STGLATKDTS CESVVTSGQH QLTPQHPPRD ASPAGLLSIA EETLTEFLSK ATGTAVEWVQ 180 MPGMKPGPDS IGIIAISHGC TGVAARACGL VGLEPSRVAE ILKDRPSWYR DCRAVDVVNV 240 LPTANGGTIE LLYMQLYAPT TLAPARDFWL LRFTSVMEDG SLVVCERSLG NVQNGPSMPP 300 VQNFVRAEML PSGYLIRPCD GGGSIIHIVD HMNVEAWSVP EVLRPLYESS AVLAQKTTMA 360 ALRQLRQIAQ EVSQTNVTNW GRRPAALRAL SHRLSRGFNE ALNGFTDEGW SLLGNDGMDD 420 VTILVNSSPD KLMGLNLSYT NGFTSISNAV LCAKASMLLQ NVPPAVLLRF LREHRSEWAD 480 NNIEAYSAAA VKVGPCSLPG ARICNFGGQV TLPLAHTIEH EEARNSLLEV IKFEGIGHSP 540 EDAMMPRDMF LLQLCSGMDE NAVGTCAELI FAPIDASFAD DAPLLPSGFR IIPLDFGKES 600 SSPNRTLDLT SALETGTAEN KVLNDLNTNN SAARSVMTIA FQFAFESHMQ ENVAAMARQY 660 VRSIISSVQR VALALSPSHL GSHGGLRLPL GTPEAHIIAR WICQSYRCYL GVELLKSTGE 720 GNESILKTLW NHSDAIVCCS AKWIVQALPV FTFANQAGLD MLETTLVALQ DITLEKIFDD 780 HGRKSLCSEF PQIMQQGFAC LQGGICMSSM GRPVSYERAV AWKVLNEEES AHCICFMFIN 840 WSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
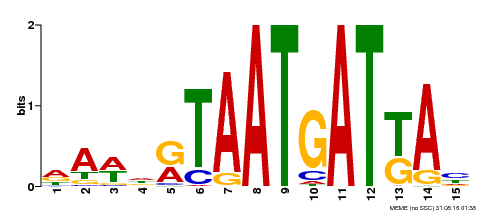
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019194412.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15-like | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A484M5A5 | 0.0 | A0A484M5A5_9ASTE; Uncharacterized protein | ||||
| STRING | XP_009593967.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA457 | 24 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



