 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000063.1_g00009.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 315aa MW: 34835.2 Da PI: 4.3293 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.1 | 7e-19 | 54 | 107 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
Itr_sc000063.1_g00009.1 54 KKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 107
56689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 131.1 | 4.3e-42 | 53 | 144 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekevee 86
ekkrrls eqvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy +Lk++yd+lk++ e+L++++e+
Itr_sc000063.1_g00009.1 53 EKKRRLSVEQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGVLKNNYDSLKHKFEALQHDNEA 138
69************************************************************************************ PP
HD-ZIP_I/II 87 Lreelk 92
L +e+
Itr_sc000063.1_g00009.1 139 LLKEIG 144
*99875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.84E-19 | 47 | 111 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.621 | 49 | 109 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.8E-18 | 52 | 113 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 4.5E-16 | 54 | 107 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.67E-17 | 54 | 110 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-20 | 56 | 116 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.8E-6 | 80 | 89 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 84 | 107 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.8E-6 | 89 | 105 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.4E-17 | 109 | 150 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 315 aa Download sequence Send to blast |
MKRSLDSSDS LGALMSICPA TTDEPSHVYS REFQTMLEGL EEEESGGGGA MTEKKRRLSV 60 EQVKALEKNF EVENKLEPER KVKLAQELGL QPRQVAVWFQ NRRARWKTKQ LERDYGVLKN 120 NYDSLKHKFE ALQHDNEALL KEIGELKAKL EGENGESKVT VKEEAVESES DNNTNNSLLL 180 RQEDETELNL EDFNGGGGLV TASLFADFKD GSSDSDSSAI LNEDNSPNAA TTGAILMISN 240 ASSSSFSSPP SINCFQFSEN PKAAAAAYQP QLVKIEEHNF FGGGDDDSCS SLFSDEQAPI 300 LHWYSPEDWN CTTSS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 101 | 109 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
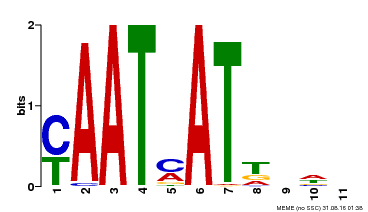
| UniProt | Transcription activator that may act as growth regulators in response to water deficit. Interacts with the core sequence 5'-CAATTATTA-3' of promoters in response to ABA and in an ABI1-dependent manner. Involved in the negative regulation of the ABA signaling pathway. {ECO:0000269|PubMed:10527431, ECO:0000269|PubMed:12065416}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By water deficit, by abscisic acid (ABA) and by salt stress. Self expression regulation. {ECO:0000269|PubMed:10527431, ECO:0000269|PubMed:12065416, ECO:0000269|PubMed:16055682}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019185794.1 | 1e-174 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like isoform X2 | ||||
| Swissprot | P46668 | 4e-73 | ATHB6_ARATH; Homeobox-leucine zipper protein ATHB-6 | ||||
| TrEMBL | A0A1S4CFT4 | 1e-123 | A0A1S4CFT4_TOBAC; homeobox-leucine zipper protein ATHB-6-like | ||||
| STRING | XP_009588928.1 | 1e-124 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2718 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 7e-69 | homeobox protein 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



