 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc000011.1_g00140.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 520aa MW: 56801.2 Da PI: 7.1044 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 51.2 | 2.3e-16 | 340 | 386 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+ellP++ K +Ka++L +A+eY+ksLq
Itr_sc000011.1_g00140.1 340 VHNLSERRRRDRINEKMRALQELLPHS-----NKTDKASMLDEAIEYLKSLQ 386
6*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 2.75E-20 | 333 | 401 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.133 | 336 | 385 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 8.53E-18 | 339 | 390 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.8E-19 | 340 | 394 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.2E-13 | 340 | 386 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.7E-17 | 342 | 391 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 520 aa Download sequence Send to blast |
MNPCLPEWNI EAEVPLANQK KPVGLDHDLV ELLWRNGQVV LHSQTHRKSG FEGNESRQLQ 60 KHDQSSLRDV SLFGNPSSFI QDDETASWLN CPVDDSFEKE FCAPFLSEIP PVHHTVGTDK 120 GGIRQVEDSK IFKVGSSEVS HSHVHSGVNL NPMPPPGTQT LSSLHQHQRR DCNFPEGGGV 180 SFGASLKANN LRSSNGQFGA KGHGDIRECS GMTVGSSHCG SNQVAIDAAF SRNSCSGNGG 240 DRGLSAAVKK DRVEKVSISP QTETAEEETA VTSSSGRSGS SFARTCNQSA GTISHGQKRK 300 VRDGEESECQ REEAAEFESG NGTKSTQKSG TPRRSRAAEV HNLSERRRRD RINEKMRALQ 360 ELLPHSNKTD KASMLDEAIE YLKSLQLQLQ MMWMGSGMAS MMFPGMQHYM SRMGMGMVPP 420 ALPNMHSAMH LPRPPMVDHA AATMASTQNQ AALCQNSMLN PINYQNQLQN PNFAEQFASY 480 MGFHPMQNPS QAVNMFNLGS HASQPNSMFN PPTNGNGPSS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 344 | 349 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
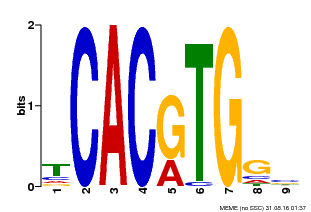
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ605441 | 0.0 | KJ605441.1 Ipomoea nil cultivar Violet PIF4 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019193873.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_019193874.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_019193875.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_019193876.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| TrEMBL | A0A0D3MTK4 | 0.0 | A0A0D3MTK4_IPONI; PIF4 | ||||
| STRING | XP_009602219.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 3e-47 | phytochrome interacting factor 4 | ||||



