 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G019050.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 452aa MW: 49887.2 Da PI: 7.0812 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.3 | 3.1e-17 | 55 | 99 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++++a k++G g W+ I +++g ++t+ q++s+ qky
HL.SW.v1.0.G019050.1 55 REKWTEEEHQKFLEALKLYGRG-WRQIEEHVG-TKTAVQIRSHAQKY 99
789*******************.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.05E-16 | 49 | 102 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 22.297 | 50 | 104 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-10 | 52 | 104 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.1E-17 | 53 | 102 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.0E-13 | 54 | 102 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.6E-15 | 55 | 99 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.12E-11 | 57 | 100 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 452 aa Download sequence Send to blast |
MAVEEQSEGM LSNNSGFAGN CIINGDANYK SESPLKDLYE SAPKVRKPYT ITKQREKWTE 60 EEHQKFLEAL KLYGRGWRQI EEHVGTKTAV QIRSHAQKYF SKVVRGSTAS VEACMKPIEI 120 PPPRPKRKPM HPYPRKSVDS FNGTSVLNQL ERSPSPNFAA TDKGTKSPTS VLSSHGSDAV 180 GYDASDHHHN RSPSSTSCTT DMQSISLSPS DKENCYLMSS SSADEKISLQ GNQLSAQSNV 240 ENFSVKIEPG CKDTARNNKE AKSSTSTSFK LFGRTVLLPG FEKESPSGTE DSETLKTDEG 300 QELDINLSLG GFVSPLESFP PINQTEQQKE SSNIVDAGAQ MLWWSMPQVP IHYQAAYNPS 360 RDQTPPKMEE KEIKKERSCS DSNEGSSSGV ETGGDRNIEA VNSFSQEPHS KGSVEQCNSR 420 KGFVPYKRCL ADRDNASTAT SDRERRKIRV H* |
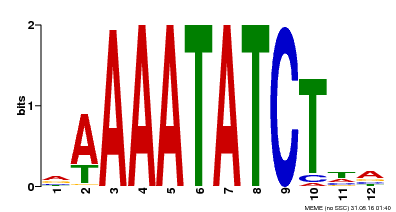
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024023521.1 | 1e-173 | protein REVEILLE 7 isoform X1 | ||||
| TrEMBL | A0A2P5CQF6 | 0.0 | A0A2P5CQF6_TREOI; GAMYB transcription factor | ||||
| STRING | XP_010099677.1 | 1e-172 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7313 | 32 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 3e-52 | MYB_related family protein | ||||



