 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.08G201900.1.p | ||||||||
| Common Name | GLYMA_08G201900, LOC100800165 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 92386.6 Da PI: 6.469 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2.1e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Glyma.08G201900.1.p 17 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 181.5 | 4.7e-57 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..E CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..g 89
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++++++l+v+ ++ g
Glyma.08G201900.1.p 161 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGLEPT-RVAEILKDRPLWFRDCRAVDVLNVLPTAngG 249
7899******************************************************.8888888888****************9999* PP
EEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SS CS
START 90 alqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgr 175
+++l +++l+a+++l+p Rdf+ +Ry+ l++g++vi+++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +
Glyma.08G201900.1.p 250 TIELLYMQLYAPTTLAPaRDFWLLRYTSVLEDGSLVICERSLKNTQNGPSmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEPW 339
************************************************9988899*********************************** PP
XXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 176 lphwllrslvksglaegaktwvatlqrqce 205
+++++lr+l++s+++ ++kt++a+l+++++
Glyma.08G201900.1.p 340 SVPEVLRPLYESSTVLAQKTTMAALRHLRQ 369
*************************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.564 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.41E-16 | 16 | 78 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.31E-16 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 5.8E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 2.37E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 25.069 | 151 | 365 | IPR002913 | START domain |
| CDD | cd08875 | 3.80E-81 | 155 | 370 | No hit | No description |
| SMART | SM00234 | 9.3E-43 | 160 | 370 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.04E-38 | 160 | 370 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 3.0E-23 | 160 | 366 | IPR023393 | START-like domain |
| Pfam | PF01852 | 1.3E-54 | 161 | 369 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.31E-5 | 397 | 491 | No hit | No description |
| SuperFamily | SSF55961 | 2.31E-5 | 521 | 596 | No hit | No description |
| Pfam | PF08670 | 3.4E-50 | 695 | 837 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MSCKDGSRNG IGMDNGKYVR YTPEQVEALE RLYHDCPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKESS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRQH 120 TQITTQATKD TNCESVVTSG QHNLTTQHPP RDASPAGLLS IAEETLAEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIVAISH GCTGVAARAC GLVGLEPTRV AEILKDRPLW FRDCRAVDVL 240 NVLPTANGGT IELLYMQLYA PTTLAPARDF WLLRYTSVLE DGSLVICERS LKNTQNGPSM 300 PPVQHFVRAE MLPSGYLIRP CEGGGSIIHI VDHMDLEPWS VPEVLRPLYE SSTVLAQKTT 360 MAALRHLRQI SHEVSQSNVT GWGRRPAALR ALSQRLSRGF NEALNGFTDE GWTTISNDGV 420 DDVTILVNSS PDKLMGLNLS FANGFPSVSN AVLCAKASML LQNVPPAILL RFLREHRSEW 480 ADNNMDAYTA AAIKVGPCSL SGSCVGNFGG QVILPLAHTI EHEEFLEVIK LEGIAHSPED 540 TIMPREMFLL QLCSGMDENA VGTCAELISA PIDASFADDA PLLPSGFRII PLESGKEASS 600 PNRTLDLASS LDVGPSGNRA SNGSAGNSSC MRSVMTIAFE FAFESHMQEH VTSMARQYVR 660 SIISSVQRVA LALSPSHLSS HAGLRSPLGT PEAQTLAHWI CNSYRCYLGV ELLKSNNEGN 720 ESLLKSLWHH SDAILCCTLK ALPVFTFSNQ AGLDMLETTL VALQDITLEK IFDDHGRKIL 780 FSEFPQIIQQ GFACLQGGIC LSSMGRPVSY ERVVAWKVLN EEENAHCICF MFVNWSFV* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.7462 | 0.0 | hypocotyl| meristem| seed coat | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed the developing vascular elements and the adaxial portion of cotyledons. Expressed in developing ovules, stamens and carpels. Expressed in procambium and shoot meristem. {ECO:0000269|PubMed:14701930, ECO:0000269|PubMed:15705957}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
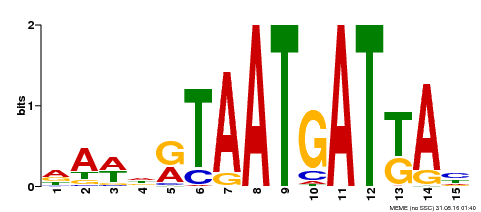
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.08G201900.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003531652.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Refseq | XP_028244497.1 | 0.0 | homeobox-leucine zipper protein ATHB-15 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A445JHH9 | 0.0 | A0A445JHH9_GLYSO; Homeobox-leucine zipper protein ATHB-15 isoform A | ||||
| TrEMBL | I1KV32 | 0.0 | I1KV32_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA08G21610.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1946 | 33 | 87 | Representative plant | OGRP651 | 16 | 71 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.1 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.08G201900.1.p |
| Entrez Gene | 100800165 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



