 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.01G022500.1.p | ||||||||
| Common Name | GLYMA_01G022500, LOC100777168 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 511aa MW: 55861.4 Da PI: 7.1975 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 47.5 | 4.3e-15 | 223 | 282 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +e+Aa+a++ a++k++g
Glyma.01G022500.1.p 223 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqaRKgRQVYLGGYDKEEKAARAYDLAALKYWG 282
57*******************666664478446*************************98 PP
| |||||||
| 2 | AP2 | 50.5 | 5.1e-16 | 325 | 376 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
Glyma.01G022500.1.p 325 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 376
57****************988532...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 7.19E-18 | 223 | 292 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 6.8E-12 | 223 | 282 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.43E-22 | 223 | 292 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.7E-16 | 224 | 291 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 4.2E-30 | 224 | 296 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.44 | 224 | 290 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.6E-6 | 225 | 236 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 6.47E-18 | 325 | 385 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.7E-10 | 325 | 376 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 9.50E-12 | 325 | 384 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.2E-18 | 326 | 384 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 6.0E-31 | 326 | 390 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.164 | 326 | 384 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.6E-6 | 366 | 386 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 511 aa Download sequence Send to blast |
MARATNWLSF SLSPMEMLRT SEPQFLQYDA ASATSSHHYY LDNLYTNGWG NGSLKFEQNL 60 NHSDVSFVES SSQSVGHVPP PPPKLEDFLG DSSAVMRYSD SQTETQDSSL THIYDHHHHH 120 HHHHGSTSYF GGDQQDLKAI TGFQAFSTNS GSEVDDSASI GKAQASEFGT HSIESSGNEF 180 AAFSGGTTGT LSLAVALSSE KAVVAAESNS SKKIVDTFGQ RTSIYRGVTR HRWTGRYEAH 240 LWDNSCRREG QARKGRQVYL GGYDKEEKAA RAYDLAALKY WGPTATTNFP VSNYSKEVEE 300 MKHVTKQEFI ASLRRKSSGF SRGASIYRGV TRHHQQGRWQ ARIGRVAGNK DLYLGTFATE 360 EEAAEAYDIA AIKFRGANAV TNFEMNRYDV EAIMKSSLPV GGAAKRLRLS LESEQKAPPV 420 NSSSQQQNPQ CGNVSGSINF SAIHQPIASI PCGIPFDSTT AYYPHNLFQH FHPTNAGAAA 480 SAVTSANATA LTALPASAAT EFFIWPHQSY * |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.27807 | 0.0 | somatic embryo | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Detected in inflorescence and youg floral mersitems, and in stem procambial cells. In floral mersitems, mostly expressed in the central dome. Disappears progressively from sepal primordia, but accumulates in second, third and fourth whorl organ primordia. Later, confined to occasional patches in stamens and in petal before disparearing progressively from flowers. {ECO:0000269|PubMed:15988559}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, seedlings, hypocotyl, inflorescence, siliques, and pistils. Also detected at low levels in leaves. {ECO:0000269|PubMed:15988559, ECO:0000269|PubMed:16307362}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
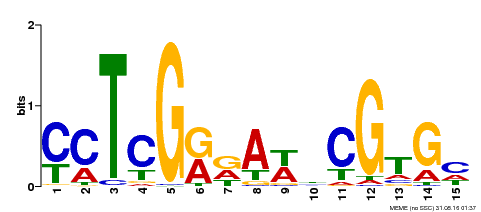
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.01G022500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015043 | 1e-102 | AP015043.1 Vigna angularis var. angularis DNA, chromosome 10, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003517351.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Refseq | XP_028228416.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A445LXS9 | 0.0 | A0A445LXS9_GLYSO; AP2-like ethylene-responsive transcription factor AIL6 isoform A | ||||
| TrEMBL | I1J503 | 0.0 | I1J503_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA01G02760.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4810 | 31 | 49 | Representative plant | OGRP112 | 17 | 209 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 1e-177 | AINTEGUMENTA-like 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.01G022500.1.p |
| Entrez Gene | 100777168 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



