 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_D02G2405 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 287aa MW: 32690.6 Da PI: 6.2428 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 56.1 | 6.4e-18 | 80 | 133 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rq+ +WFqNrRa++k
Gh_D02G2405 80 KKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWK 133
456899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 131.8 | 2.7e-42 | 79 | 171 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl+ eqvk+LE++Fe +kLeperK++lar+Lglqprq+a+WFqnrRAR+ktkqlEkdy+ Lkr+y+a+k++n++L++++++L++e+++
Gh_D02G2405 79 EKKRRLNMEQVKTLEKNFELGNKLEPERKMQLARALGLQPRQIAIWFQNRRARWKTKQLEKDYDLLKRQYEAIKADNDALQTQNQKLHAEIMA 171
69**************************************************************************************99975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.71E-19 | 70 | 137 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.329 | 75 | 135 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.8E-17 | 78 | 139 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.49E-16 | 80 | 136 | No hit | No description |
| Pfam | PF00046 | 2.9E-15 | 80 | 133 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 4.7E-19 | 82 | 142 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 1.1E-5 | 106 | 115 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 110 | 133 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.1E-5 | 115 | 131 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.1E-16 | 135 | 175 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 287 aa Download sequence Send to blast |
MSCNGMAFFP ANFMLQSPHE EDHQPSTSIN QMLPSCPPQD FHGVASFLGK RSMSFSGIDV 60 CEEANGEDDL SDDGSQAGEK KRRLNMEQVK TLEKNFELGN KLEPERKMQL ARALGLQPRQ 120 IAIWFQNRRA RWKTKQLEKD YDLLKRQYEA IKADNDALQT QNQKLHAEIM ALKSREPTES 180 INLNKETEGS CSNRSENSSD VKLDISRTPA IDSPLSTHPT SRTFFPTSIR PTGGAAQLFQ 240 TSSSRPDHHH PCQKMDQMVK EESLSNMFCT IEDQTGFWPW LEQPHFN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 127 | 135 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ghi.11309 | 0.0 | boll | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Predominantly expressed in leaves and flowers. {ECO:0000269|PubMed:16055682}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may act in the sucrose-signaling pathway. {ECO:0000269|PubMed:11292072}. | |||||
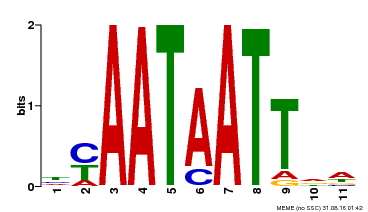
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012478668.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-13 | ||||
| Refseq | XP_012478669.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-13 | ||||
| Swissprot | Q8LC03 | 1e-121 | ATB13_ARATH; Homeobox-leucine zipper protein ATHB-13 | ||||
| TrEMBL | A0A0D2SGM9 | 0.0 | A0A0D2SGM9_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.005G139600.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2633 | 27 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 1e-123 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



