 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Genemark1.1008_g | ||||||||
| Common Name | COCSUDRAFT_59046 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Trebouxiophyceae incertae sedis; Coccomyxaceae; Coccomyxa; Coccomyxa subellipsoidea
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 575aa MW: 60565.6 Da PI: 6.1371 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 35.4 | 2.4e-11 | 17 | 52 | 1 | 38 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlk 38
r +W+ E++++ +av+++G + Wk I +++g +R+
Genemark1.1008_g 17 REKWSDSEHQRFTEAVEKYGRD-WKMIVEHVG-TRSVA 52
789*****************77.*********.**975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.09E-9 | 8 | 53 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 10.696 | 12 | 85 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 7.1E-8 | 15 | 52 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 0.0058 | 16 | 58 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-6 | 16 | 51 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 7.6E-9 | 17 | 53 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.30E-7 | 19 | 53 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 575 aa Download sequence Send to blast |
MADAPPTRKP YRITKQREKW SDSEHQRFTE AVEKYGRDWK MIVEHVGTRS VAQSSLGQLR 60 DDVLPSCGGP VPVRSHAQKF FLKLEKSGQA GVVPPPRPKK RAAKPYPVQD RGEERRKSKR 120 SRSSASNSSA HSQPSQTRGA AVSPSTSPSN GRSSPRSAAT AATLPVSSRR PLRSSRATRS 180 LANAAASTKG SSQDLEGNGS NIASANESNN DHSEGPSATQ ENSGLSTGGA DLDLQFSADP 240 MSLEAPEAAP RPREKSETPD YGKLYSIVNV LFDSGGAAND EKNHKQMMEQ LEPGQRKELL 300 AMMDQLTGNL RSTCPPAQKL EQLKAELKAK SQSPTPPQQP PPTPIPAPAR ASRLDHAASM 360 PLMPCTACLP MPQTVSEEPL ALMQPLPGSQ GTALLQPLPC AQQCSFSPVL PPPLASDACE 420 LVHCGSGAGS SLGSGIGCGS VAALPGLAAD AVLLQQSEPE QSIAEQLLVD NSGEGAGMGQ 480 AMFDIQQGSG LEEVGLQLPK SSSLGGNTPL DTCLAQLESG TDPTVPESDD FLDSWKIGSS 540 DPVWQTLEMQ QLSPTAPTSP SSFTRDTNGS MFAL* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
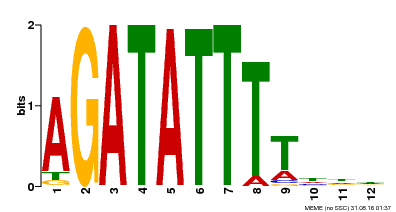
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005651061.1 | 0.0 | hypothetical protein COCSUDRAFT_59046 | ||||
| TrEMBL | I0Z798 | 0.0 | I0Z798_COCSC; Uncharacterized protein | ||||
| STRING | XP_005651061.1 | 0.0 | (Coccomyxa subellipsoidea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP454 | 14 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G37260.1 | 1e-19 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17044527 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



