 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01034505001 | ||||||||
| Common Name | LOC100248677, VIT_18s0072g00840, VITISV_010700 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 263aa MW: 28654.9 Da PI: 6.1244 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 20.6 | 1.2e-06 | 41 | 62 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
C +Cgk F+r nL+ H+r H
GSVIVT01034505001 41 FCMICGKGFKRDANLRMHMRGH 62
6*******************98 PP
| |||||||
| 2 | zf-C2H2 | 11 | 0.0013 | 127 | 142 | 6 | 21 |
TTEEESSHHHHHHHHH CS
zf-C2H2 6 CgksFsrksnLkrHir 21
Cg++Fsrk+ L Hi
GSVIVT01034505001 127 CGTTFSRKDKLFGHIA 142
**************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 1.21E-5 | 39 | 62 | No hit | No description |
| SMART | SM00355 | 0.0057 | 40 | 62 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.697 | 40 | 67 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 41 | 66 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 42 | 62 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.21E-5 | 90 | 116 | No hit | No description |
| SMART | SM00355 | 22 | 95 | 117 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010044 | Biological Process | response to aluminum ion | ||||
| GO:0010447 | Biological Process | response to acidic pH | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MNGEHELKDE DDADEGENLP PGSYEILQLE KEEILAPHTH FCMICGKGFK RDANLRMHMR 60 GHGDEYKTPA ALAKPNKESS SEPVLIKRTH CDKSYTCSRC NTKKFSVIAD LKTHEKHCGK 120 DKWLCSCGTT FSRKDKLFGH IALFQGHTPA IPLDETKGSV GPSDRGEGNG AANKVGSVGF 180 NFSSNASSGS GVQDMMMDVK RGADEPTGFF SPLTFDPCSL VGFHEFPRPP FEDSESSFSF 240 LVPGSCSYTR KTGGESSSND LE* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.9482 | 0.0 | bud| cell culture| flower| fruit| inflorescence| leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots (e.g. root tips and lateral roots), leaves, flowers (e.g. stigma, sepal, anther, and filament), stems, siliques and cotyledons. {ECO:0000269|PubMed:23935008}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Together with STOP2, plays a critical role in tolerance to major stress factors in acid soils such as proton H(+) and aluminum ion Al(3+). Required for the expression of genes in response to acidic stress (e.g. ALMT1 and MATE), and Al-activated citrate exudation. {ECO:0000269|PubMed:17535918, ECO:0000269|PubMed:18826429, ECO:0000269|PubMed:19321711, ECO:0000269|PubMed:23935008}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00196 | ampDAP | Transfer from AT1G34370 | Download |
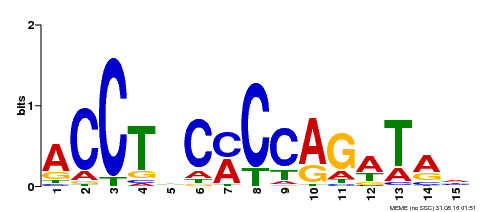
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By shock H(+) and Al(3+) treatments. {ECO:0000269|PubMed:17535918}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM469305 | 0.0 | AM469305.2 Vitis vinifera contig VV78X267210.11, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019072266.1 | 0.0 | PREDICTED: protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| Refseq | XP_019072267.1 | 0.0 | PREDICTED: protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| Swissprot | Q9C8N5 | 1e-119 | STOP1_ARATH; Protein SENSITIVE TO PROTON RHIZOTOXICITY 1 | ||||
| TrEMBL | A5BSF5 | 0.0 | A5BSF5_VITVI; Uncharacterized protein | ||||
| STRING | VIT_18s0072g00840.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP830 | 16 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34370.3 | 1e-105 | C2H2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01034505001 |
| Entrez Gene | 100248677 |



