 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01029593001 | ||||||||
| Common Name | LOC100251358, VIT_09s0054g00440 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 245aa MW: 26666.9 Da PI: 8.6425 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 51.5 | 1.4e-16 | 165 | 201 | 1 | 35 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkgl 35
C++Cg ++ Tp++Rrgp+g++tLCnaCGl+++ kg+
GSVIVT01029593001 165 CRHCGISEksTPMMRRGPEGPRTLCNACGLMWANKGT 201
*****99999************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00979 | 8.9E-12 | 25 | 60 | IPR010399 | Tify domain |
| PROSITE profile | PS51320 | 14.846 | 25 | 60 | IPR010399 | Tify domain |
| Pfam | PF06200 | 1.2E-13 | 28 | 59 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 13.205 | 91 | 133 | IPR010402 | CCT domain |
| Pfam | PF06203 | 4.6E-16 | 91 | 133 | IPR010402 | CCT domain |
| SMART | SM00401 | 3.7E-13 | 159 | 212 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 8.748 | 159 | 207 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 1.14E-12 | 161 | 210 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 4.8E-16 | 163 | 206 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 4.41E-15 | 164 | 219 | No hit | No description |
| Pfam | PF00320 | 1.5E-14 | 165 | 201 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 165 | 192 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045944 | Biological Process | positive regulation of transcription from RNA polymerase II promoter | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0000977 | Molecular Function | RNA polymerase II regulatory region sequence-specific DNA binding | ||||
| GO:0001085 | Molecular Function | RNA polymerase II transcription factor binding | ||||
| GO:0001228 | Molecular Function | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 245 aa Download sequence Send to blast |
MEGDVQADPG NLADSRGALT VQPGGSDNQN QLTLSFQGQV YVFDSVSPEK VQAVLLLLGG 60 REVPPTMPAL SIAGHNRELP GTPQRYNVPH RLASLIRFRE KRKERNFDKK IRYTVRKEVA 120 LRMQRNKGQF TSSKSNHDDS ASTTPGWESS WGMAGNGPIN QEIVCRHCGI SEKSTPMMRR 180 GPEGPRTLCN ACGLMWANKG TLRDLSKAAP EAGQSPSLNQ TGENGNFETD QMVLGMAENV 240 SDQA* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.12431 | 0.0 | fruit | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Predominantly expressed in shoot apices, inflorescences and roots. {ECO:0000269|PubMed:14966217}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00198 | DAP | Transfer from AT1G51600 | Download |
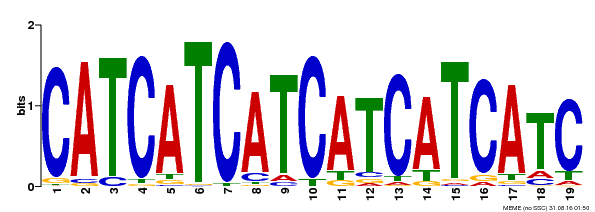
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FQ393169 | 0.0 | FQ393169.1 Vitis vinifera clone SS0AFA2YF05. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002263707.1 | 0.0 | PREDICTED: GATA transcription factor 24 | ||||
| Refseq | XP_010655300.1 | 0.0 | PREDICTED: GATA transcription factor 24 | ||||
| Refseq | XP_010655302.1 | 0.0 | PREDICTED: GATA transcription factor 24 | ||||
| Swissprot | Q8H1G0 | 1e-104 | GAT28_ARATH; GATA transcription factor 28 | ||||
| TrEMBL | F6GYN7 | 0.0 | F6GYN7_VITVI; Uncharacterized protein | ||||
| STRING | VIT_09s0054g00440.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP831 | 16 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G51600.2 | 1e-107 | ZIM-LIKE 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01029593001 |
| Entrez Gene | 100251358 |



