 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01029265001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 280aa MW: 30419.6 Da PI: 10.4421 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 100 | 1.4e-31 | 207 | 265 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-ST.T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsa.gCpvkkkversaedpkvveitYegeHnhe 59
D+y+WrKYGqK++kgs++pr+YY+C+s gCp++k+ver+ +dp ++++tYegeH+h+
GSVIVT01029265001 207 ADEYSWRKYGQKPIKGSPYPRGYYKCSSVrGCPARKHVERAPDDPAMLIVTYEGEHRHS 265
69*************************988****************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF10533 | 2.2E-20 | 157 | 204 | IPR018872 | Zn-cluster domain |
| Gene3D | G3DSA:2.20.25.80 | 2.4E-33 | 194 | 266 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 30.148 | 201 | 267 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.57E-26 | 204 | 266 | IPR003657 | WRKY domain |
| SMART | SM00774 | 5.8E-38 | 206 | 266 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 6.2E-27 | 208 | 264 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 280 aa Download sequence Send to blast |
MAVDFLGFSK MDEQMAIQEA ASAGLKSMEH LILLLNHHHP QSQQINHFDC REITDFTVSK 60 FKQVISILNR TGHARFRRGP VTSSPSHPFH SNLILSAKPD PLKSEGNASV SSTTSSFLSS 120 ITGDGSVSNG KLGTPLFAPP PAPAVSAGKP PLSSSQRRKC HEHGSSDNIS GKLSVSGRCH 180 CSKRRKNRVK RTIRVPAISS KIADIPADEY SWRKYGQKPI KGSPYPRGYY KCSSVRGCPA 240 RKHVERAPDD PAMLIVTYEG EHRHSQTPAP AGGLMFPST* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 2e-22 | 194 | 269 | 2 | 76 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.11310 | 0.0 | bud| flower| inflorescence| leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: In young, mature and senescent leaves. {ECO:0000269|PubMed:11722756}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). Regulates rhizobacterium B.cereus AR156-induced systemic resistance (ISR) to P.syringae pv. tomato DC3000, probably by activating the jasmonic acid (JA)- signaling pathway (PubMed:26433201). {ECO:0000250, ECO:0000269|PubMed:26433201}. | |||||
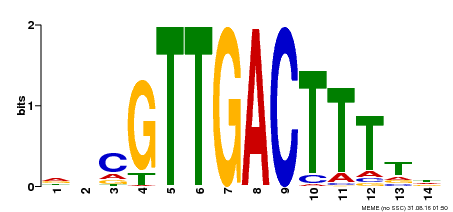
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00464 | DAP | Transfer from AT4G31550 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Strongly during leaf senescence (PubMed:11722756). Repressed by rhizobacterium B.cereus AR156 in leaves, and to a lower extent, by P.fluorescens WCS417r (PubMed:26433201). {ECO:0000269|PubMed:11722756, ECO:0000269|PubMed:26433201}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU056837 | 1e-159 | GU056837.1 Vitis vinifera WRKY gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002266188.1 | 1e-175 | PREDICTED: WRKY transcription factor WRKY51 | ||||
| Swissprot | Q9SV15 | 1e-108 | WRK11_ARATH; Probable WRKY transcription factor 11 | ||||
| TrEMBL | D0V9L3 | 1e-173 | D0V9L3_VITVI; WRKY | ||||
| STRING | XP_010265415.1 | 1e-132 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP3557 | 11 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G31550.1 | 7e-70 | WRKY DNA-binding protein 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01029265001 |



