 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01011942001 | ||||||||
| Common Name | VIT_01s0011g03110 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 357aa MW: 39255.4 Da PI: 8.1641 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.4 | 2.9e-32 | 188 | 243 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs++AtPk+i+elmkv+gLt+++vkSHLQkYRl+
GSVIVT01011942001 188 KARRCWSPELHRRFLHALQQLGGSHVATPKQIRELMKVDGLTNDEVKSHLQKYRLH 243
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.219 | 185 | 245 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-28 | 185 | 246 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.54E-18 | 186 | 246 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.7E-27 | 188 | 243 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 190 | 241 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 357 aa Download sequence Send to blast |
MQRCHDYIEA LEEERRKIQV FQRELPLCLE LVSQAIESCR QQMSGTTQEY FHGQSECSEQ 60 TSSDGPVLEE FIPIKKTSDD EDEQQSHQPN DNKDKNNDKS GKKSDWLRSV QLWNQTPDPP 120 VKEDTPKKIP SMEVKKNGGA FHPFKRDKAV GTNPTSAPSA ATSSTAETAT GCSSGSRKEE 180 KEGQSQRKAR RCWSPELHRR FLHALQQLGG SHVATPKQIR ELMKVDGLTN DEVKSHLQKY 240 RLHTRRPNPA IQHNGNPQAP QFVVVGGIWV PPPEYTAVAA TTSSGEATGV TTANGIYAPV 300 ASVPPSHPQG STQRQQPMKP KKSQSEERGS HSEGGVQSNS PATSSSTHTT TTSPVF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-13 | 188 | 241 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-13 | 188 | 241 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-13 | 188 | 241 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-13 | 188 | 241 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-13 | 188 | 241 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
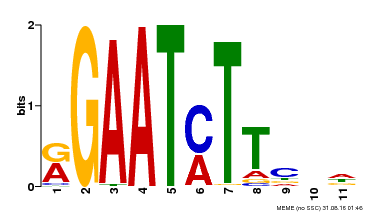
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FQ383630 | 0.0 | FQ383630.1 Vitis vinifera clone SS0ABG1YE07. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003631224.1 | 0.0 | PREDICTED: myb family transcription factor EFM | ||||
| Swissprot | Q9FPE8 | 1e-109 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A438ECN5 | 0.0 | A0A438ECN5_VITVI; Transcription factor HHO3 | ||||
| STRING | VIT_01s0011g03110.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP5415 | 12 | 21 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 1e-101 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01011942001 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



