 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSVIVT01009845001 | ||||||||
| Common Name | LOC100233070, VIT_18s0001g13730 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; rosids incertae sedis; Vitales; Vitaceae; Vitis
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 147aa MW: 16107.6 Da PI: 10.4186 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 274 | 7.5e-84 | 5 | 146 | 158 | 301 |
GAGA_bind 158 kpkekkakkkkkksekskkkvkkesaderskaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyPLPvstk 249
k+ +k kk+k+++e+ +k v +s sk+++k +dl+ln+v++Dest+P+PvCsCtG+lrqCYkWGnGGWqS+CCttt+S+yPLP++++
GSVIVT01009845001 5 KKASKPLKKPKREGEDLNKIVFGKS--LVSKSDWKGQDLGLNQVTFDESTMPAPVCSCTGVLRQCYKWGNGGWQSSCCTTTLSMYPLPAVPN 94
2233334566666666665555555..67*************************************************************** PP
GAGA_bind 250 rrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
+r+aR++grKmS++af+klL++LaaeG+dls pvDLkdhWAkHGtn+++ti+
GSVIVT01009845001 95 KRHARVGGRKMSGSAFNKLLSRLAAEGHDLSIPVDLKDHWAKHGTNRYITIK 146
***************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 2.9E-62 | 1 | 146 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 2.4E-82 | 4 | 146 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 147 aa Download sequence Send to blast |
MSSNKKASKP LKKPKREGED LNKIVFGKSL VSKSDWKGQD LGLNQVTFDE STMPAPVCSC 60 TGVLRQCYKW GNGGWQSSCC TTTLSMYPLP AVPNKRHARV GGRKMSGSAF NKLLSRLAAE 120 GHDLSIPVDL KDHWAKHGTN RYITIK* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 4 | 12 | KKASKPLKK |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Vvi.3407 | 0.0 | bud| cell culture| flower| fruit| inflorescence| pedicel | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in seedlings, leaves and pistils. Detected in the base of flowers and tips of carpels, in sepal vasculature, in young rosette, in the lateral and tip of primary roots, and in ovule at the exception of the outer integument. {ECO:0000269|PubMed:14731261, ECO:0000269|PubMed:21435046}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
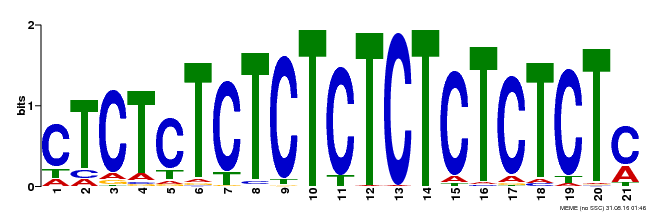
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU597746 | 0.0 | EU597746.1 Vitis vinifera GAGA-binding transcriptional activator BBR/BPC6-like gene, complete cds. | |||
| GenBank | EU597747 | 0.0 | EU597747.1 Vitis vinifera GAGA-binding transcriptional activator BBR/BPC6-like mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001268020.1 | 1e-100 | GAGA-binding transcriptional activator BBR/BPC6-like | ||||
| Refseq | XP_010664073.1 | 1e-100 | PREDICTED: GAGA-binding transcriptional activator BBR/BPC6-like isoform X1 | ||||
| Swissprot | Q8L999 | 4e-79 | BPC6_ARATH; Protein BASIC PENTACYSTEINE6 | ||||
| TrEMBL | B2Z458 | 3e-99 | B2Z458_VITVI; GAGA-binding transcriptional activator BBR/BPC6-like | ||||
| STRING | VIT_18s0001g13730.t01 | 1e-100 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP2841 | 13 | 31 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.3 | 7e-79 | basic pentacysteine 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | GSVIVT01009845001 |
| Entrez Gene | 100233070 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



